

| August 27, 2004 | science beat | | lab a-z index | lab home |
 |
 |
||
|
|||
| A Push-Pull Approach to Proteins Researchers learn the biophysical properties of bacterial condensin |
| Contact : Dan Krotz, dakrotz@lbl.gov |
|
By stretching a poorly understood protein like a rubber band, a team of Berkeley Lab and University of California at Berkeley scientists is learning how the protein and its cousins perform some of life's most fundamental tasks. Their work, published in the journal Science, is the first look at the biophysical properties of a condensin. Condensins help in the final stages of folding coils of DNA into chromosomes. They are found in all organisms, from bacteria to people, and are part of a family of proteins responsible for other critical jobs such as DNA repair and meiosis. Despite its importance, condensin was discovered only ten years ago and scientists are only now beginning to understand how it works. "If we can understand how condensin works, it will help us understand how a large group of very important proteins work," says Carlos Bustamante, a biophysicist with Berkeley Lab's Physical Biosciences Division and UC Berkeley's Department of Molecular and Cell Biology, who is also a Howard Hughes Medical Institute investigator. "But we couldn't study the protein in a bulk assay as is normally done, because such an assay doesn't exist for condensin. So we took a risk." 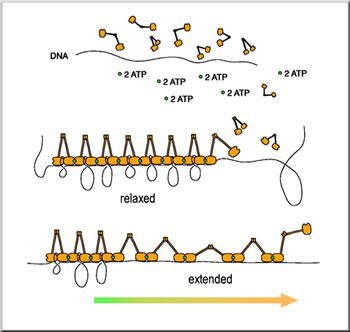
Pincer-like MukBEF complexes trap and supercoil lengths of nucleosomal DNA between their heads (orange), with each pincer joined to the next by ATP (green). When the filament is extended the heads break apart to release varying lengths of DNA, but the MukBEF pincers remain attached to one another by ATP. When force on the DNA is relaxed, MuKBEF pincers close and recoil the DNA. Instead, they analyzed its structure by stretching a single molecule of DNA, which had been condensed into a fiber by the binding of many condensin molecules, until some of the bonds within the condensin molecules snapped. This effect, like pulling a series of slipknots out of a rope, enabled the team to measure the strength of the broken bonds and infer how the condensin molecules are pieced together in the fiber. "We were able to study, very clearly, how each one of the condensin molecules bound to the DNA," says Bustamante. "And this gives us hope that we can decipher how a large family of proteins interacts with DNA." Bustamante conducted the research with Nicholas Cozzarelli of UC Berkeley's Department of Molecular and Cell Biology and Steve Smith of the Howard Hughes Medical Institute. Bustamante is also part of an interdisciplinary Physical Biosciences Division initiative called Microscopies of Molecular Machines (M3) that uses conventional and newly developed microscopy techniques to probe the inner workings of molecular machines. Earlier protein crystallography and electron microscopy research had revealed that a single condensin is about 100 nanometers long (one nanometer is one-billionth of a meter) and shaped like a nutcracker, with two rods ending at a globular domain or head, joined by a central hinge. In addition, genetic and biochemical experiments had revealed that the proteins are closely involved in maintaining the compact organization of DNA inside cells. Although this work went a long way in describing condensin's biochemical makeup, scientists didn't understand the protein's mechanical properties, or how it helps engineer one of nature's greatest packing jobs. In DNA folding, DNA is rolled into nucleosomes, nucleosomes are coiled into 30-nanometer wide fibers, and these fibers are attached to condensins, which oversee the final compacting and folding necessary to squeeze the fibers into chromosomes. This last step puts the finishing touches on a process that somehow compresses about one meter of DNA into every human cell. To find out how condensin helps facilitate this feat, the research team allowed the Escherichia coli condensin, called MukBEF, to bind to a single DNA molecule strung between two microscopic beads. In the presence of ATP, the condensin molecules greatly reduced the dimensions of the DNA molecule by condensing it in an ATP-dependent manner. Next, the researchers slowly stretched this condensed fiber, gradually subjecting the bonds holding the proteins together to greater and greater stress. At 17 piconewtons (one newton is approximately the weight of one apple, and one piconewton is the weight of one red blood cell), they observed a rapid-fire series of ruptures as the intramolecular contacts between the adjacent heads of a condensin molecule broke apart. At the same time, the intermolecular interactions between the heads of adjacent condensin molecules on the DNA, and their bonds to the DNA, remained intact. In addition, when the researchers reduced the stress on the fiber by moving the two beads closer together, the protein accordioned back to its original shape, even if the excess free ATP in solution had been removed. In other words, after being deformed microns in length, the filament returns to its compact structure without the addition of (ATP) energy. 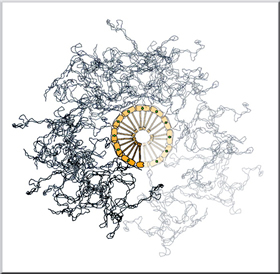
Loops of supercoiled nucleosomal DNA form domains between heads of MukBEF complexes, which are thought to propagate so as to form a helix. "We were able to directly probe the forces that maintain the structure of the protein assembly on the DNA," says Bustamante. "And from this mechanistic analysis, we could infer the way in which the protein assembles itself into a larger complex." Based on these results, Bustamante, Cozzarelli, and Smith conclude that MukBEF forms a regularly ordered filament linear DNA in which low-energy interactions are broken and high-energy attachments survive. These properties may also help explain how close relatives of condensin maintain the genome through jobs such as homologous DNA repair, in which broken DNA ends are repaired using information from intact DNA. Perhaps the contortionist-like protein pulls the broken and unbroken DNA strands closer together so they can exchange information. Although more work needs to be conducted on this family of proteins, such as measuring the precise energy involved in the molecular interactions holding them together, this initial inquiry into a condensin's biophysical traits sheds light on the inner-workings of life. "Crystallography tells you how a protein looks, genetics tells you what it does, biochemistry tells you what it interacts with, and biophysics tells you how it works," says Ryan Case of Berkeley Lab's Materials Sciences Division and UC Berkeley's Department of Molecular and Cell Biology, a postdoctoral researcher in the Bustamante laboratory who, with graduate students Yun Pei Chang and Jeff Gore in the same lab, performed most of the experiments. "If you want to know how a car works, you take it apart, you don't just look at it." This work was supported in part by grants from the Department of Energy's Office of Biological and Environmental Research and the National Institutes of Health. Additional information
|
| Top |