|
The ties that bind
To repair a double-strand break, the ends of the broken strands must be
stripped of damaged or redundant bases, then brought together with a new
length of DNA. The new length may be assembled intact from a nearby "sister"
chromatid -- the adjacent arm of the chromosome -- or it may involve an
imperfectly matched length of DNA in a process called nonhomologous end-joining.
In both processes, the preparatory work on a broken strand is done by
a complex of two Mre11s and two Rad50s that form a "binding head."
The binding head is compact, but each of its Rad50s has a long tail,
extending up to 600 angstroms in mammals, which consists of a coil of
amino acids coiled back around itself -- like a twisted fiber pinched
in the middle, its two halves wound around each other to form a loose
piece of twine.
|
 |
 |
|
|
 |
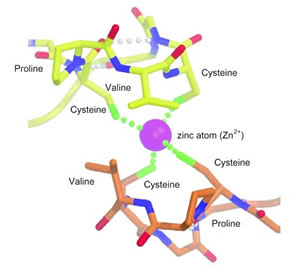
Rad50 tails link together when four
cysteine amino-acid residues in the CXXC motifs (two cysteines each,
in a pair of Rad50 tails) bind to a single, doubly ionized zinc atom.
|
|
At the pinch in the middle of the primary coil, where the tail bends
back on itself, there is a sequence of four amino acid residues similar
in all the organisms studied. The first and fourth residues in this sequence
are always cysteines, with various other residues in the second and third
places -- thus the label "CXXC motif."
Through x-ray crystallography the researchers learned that the CXXC motifs
were shaped like hooks, with the potential to grapple the tails of other
Rad50 proteins. Using laboratory techniques of gel filtration and ultracentrifugation,
the group confirmed that successful linkage of two Rad50 proteins depends
on the presence of zinc.
Chemical and structural studies showed that four cysteines (two each,
in a pair of Rad50 tails) bind to a single, doubly ionized zinc atom.
Discovery of this zinc-hook mechanism immediately suggested ways the Mre11
complex could connect DNA strands and bring them together.
A biological multitool kit
In a series of compelling electron micrographs of human and archaean Mre11
complexes, the researchers found several different conformations of linked
Rad50. Some pairs were joined by their tail hooks to form a bola shape,
with the binding heads of the Mre11 complex at both ends.
In one micrograph, the twin tails of a single binding head link to each
other, forming what looks like a finger ring, with the binding head as
its stone; an even more unusual micrograph shows a single-headed complex
of this kind actually bound to a length of DNA.
These conformations suggest distinct mechanisms by which linked Mre11
complexes can bridge sister chromatids or perform nonhomologous end-joining,
and how even a circular, single-headed Mre11 could bring together two
broken DNA ends.
 |
|
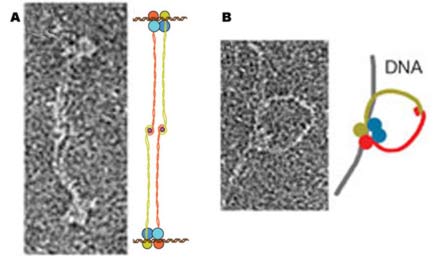
| Electron micrographs show two possible
structures formed by Rad50 zinc hooks: (A) two Mre11/Rad50 complexes
could link their tails and attach to separate double strands of DNA
with their binding heads; (B) a single Mre11/Rad50 complex, its twin
tails linked in a circle, joins a broken double strand. |
| |
"Identification of the Rad50 zinc hook will allow us to understand
the mechanism for dynamic assembly and disassembly of the Mre11 complex,"
says Tainer, "which is critical for the repair of double-strand breaks
in DNA and thus for the avoidance of cancer-causing mutations."
The discovery is exciting not only for the questions it answers but for
the new questions it raises. Do binding heads begin their work separately,
thrashing their tails about until they grapple another bound complex that's
likely to help complete the repair? Do they link tails first and search
together for broken DNA to repair?
Other models are possible; which are correct and how the process works
in detail are yet to be determined, but other coordinating proteins and
interactions are likely to be involved. Thus finding the correct model
requires the complete NCI-funded team.
SBDR's leaders emphasize the cyclical nature of their work, first drawing
on genome sequencing and biochemical and genetic studies to suggest the
proteins most worth studying, then determining the structures of these
proteins through x-ray crystallography and other methods, in search of
understanding of how they work together to accomplish complex biological
tasks.
This knowledge, in turn, suggests biological experiments that can give
clues to the make-up and mechanisms underlying the complex machines that
mediate DNA repair, filling in more details of their intricate interactions
-- a new kind of research design cycle that provides a novel prototype
for productively linking structure and chemistry with biology.
Additional information:
|