

Lab BioSciences Get Boost From Million-Dollar Grants
Blast from the Past Through Neanderthal DNA Sequencing
Lab BioSciences Get Boost From Million-Dollar Grants
Research at Berkeley Lab aimed at furthering the understanding of genomic and proteomic aspects of cancer, and increasing the use of crystallography to identify the structures of proteins and other macromolecules, received a welcome boost this fall from several new multi-million dollar grants.
 Paul Adams, an authority on macromolecular crystallography, heads the Berkeley Center for Structural Biology and is a senior scientist in the Physical Biosciences Division. He is shown here at ALS beamline 8.2.2, a superbend beamline funded by HHMI, with Corie Ralston, who co-manages the beamline with Adams.
Paul Adams, an authority on macromolecular crystallography, heads the Berkeley Center for Structural Biology and is a senior scientist in the Physical Biosciences Division. He is shown here at ALS beamline 8.2.2, a superbend beamline funded by HHMI, with Corie Ralston, who co-manages the beamline with Adams.
The autumn windfalls started in September when the National Cancer Institute (NCI) announced awards totaling over $35.5 million for the establishment of regional teams to find cancer-specific proteins and protein patterns in the blood that can be used to detect cancer. Researchers with the Lab’s Life Sciences Division will be members of the team based in the San Francisco Bay Area, in partnership with UC San Francisco and the Buck Institute for Age Research.
In early October, NCI and the National Human Genome Research Institute (NHGRI) named the newly established Berkeley Cancer Genome Center in the Life Sciences Division as one of seven programs that will share $35 million over the next three years as part of an effort to identify important genetic changes involved in lung, brain, and ovarian cancers.
Later that same month, the National Institutes of Health (NIH) awarded an $8.2 million grant to researchers in the Physical Biosci-ences Division to develop the next generation of a software program called PHENIX, which automates crystallography data acquisition and analysis. In addition, a $4.8 million pledge was received from the Howard Hughes Medical Institute (HHMI) to upgrade the robotic capabilities of their beamlines at the Advanced Light Source.
Proteomics is the study of all the proteins in a cell, tissue, or organism, called its proteome, just as all of an organism’s genes are called its genome. The goal of clinical proteomics for early detection of cancer is to identify certain proteins or patterns of proteins in bodily fluids, such as blood serum, which may signal cancer long before current methods of diagnosis reveal cancer in a patient.
NCI, which is a part of NIH, launched its Clinical Proteomic Assessment for Cancer (CPTAC) initiative with the goal of identifying new protein biomarkers that can be tested for, as is now done with the prostate specific antigen test for prostate cancer. Leading the Bay Area’s CPTAC team is Susan Fisher, who holds a joint appointment with the Life Sciences Division and UCSF. Joe Gray, director of the Life Sciences Division, is one of the team’s co-principal investigators.
“Our team’s approach will be to combine information from genome-wide studies of gene expression and gene processing with mass-spectrometry-based protein analysis technologies to identify cancer-specific proteins in the blood,” Gray said.
Gray also co-directs, with Life Sciences’ Paul Spellman, the Berkeley Cancer Genome Center, which is a collaboration between Berkeley Lab, UCSF and UC Berkeley. With the new grants from NCI and NHGRI, researchers at the center will focus on identifying changes to the populations of messenger RNA that occur in cancer. Such changes are indicative of different kinds of proteins produced by the altered genomes of tumor cells.
Said Spellman, “The Center will use the Affymetric Exon 1.0 array platform to measure exon-specific expression (exons are the coding sequences in a gene) of at least 1,000 samples per year, and will use computational tools to identify those whose behavior suggests they might play a role in cancer.”

Joe Gray
With genome sequencing having reached nearly a mass-production mode and new genes being identified on a regular basis, there is a growing demand for faster methods of identifying the structures of the proteins and nucleic acids being produced by these genes. The PHENIX software provides algorithms for high-throughput protein structure determination via x-ray crystallography. PHENIX stands for Python-based Hierarchical Environment for Integrated Xtallography, and it was originally developed through a multi-institutional collaboration led by Paul Adams, of Berkeley Lab’s Physical Biosciences Division.
Under the grant from NIH’s National Institute for General Medical Sciences, Adams and a new collaboration will spend the next five years developing algorithms that will extend PHENIX extend beyond the realm of structural genomics and allow crystallographers to solve other challenging biological problems as well, including modeling nucleic acid molecules.
“DNA and RNA represent a very important class of molecule crucial to understanding biology, yet even though more than 1,000 structures containing nucleic acids are in the Worldwide Protein Data Bank, there are no automated procedures for building models of these molecules,” said Adams. “We will fill this gap by developing methods to automatically build, refine and validate nucleic acid structures.”
Adams is the director of the Berkeley Center for Structural Biology (BCSB) which operates five macromolecular crystallography beamlines at the ALS, including beamlines 8.2.1 and 8.2.2, two “superbend” beamlines which were built through funding by HHMI. With the latest grant from HHMI, the BCSB plans to equip beamlines 8.2.1 and 8.2.2 with robotic crystal auto-mounting equipment that will enable researchers to take advantage of high throughput crystal screening. Both beamlines will also receive new and improved x-ray fluorescence detectors that will permit the detection of weaker signals from small or dilute samples, and both will also receive upgraded computer hardware.
In addition, the end station at beamline 8.2.1 will be provided with an upgraded large-format CCD detector, to facilitate high-resolution data collection and the study of crystals with large unit cells, and beamline 8.2.2 will receive an upgrade of its optics to decrease the size of its x-ray beam so that diffraction experiments can be carried out on crystals as tiny as 10 microns in diameter, about the size of a human cell.
Said Adams, “Crystallographic structure solution for a wealth of proteins and other macromolecules has often been hindered by the small size of available crystals. The ability to focus a high-brightness x-ray beam into a size small enough to match the dimensions of a crystal should allow us to make optimal scientific use of that crystal.”
Blast from the Past Through Neanderthal DNA Sequencing
“While we are not going to be able to bring Neanderthals back to life, their DNA sequences can serve as a time machine enabling us to go back and learn about the biology of these extinct hominids in ways we haven’t been able to do in the 150 years since the first Neanderthal was unearthed.”
 Neanderthals are the closest hominid relatives of modern humans. The two species co-existed in Europe and western Asia as late as 30,000 years ago.
Neanderthals are the closest hominid relatives of modern humans. The two species co-existed in Europe and western Asia as late as 30,000 years ago.
So said Berkeley Lab research geneticist and director of the Joint Genome Institute, Edward (Eddy) Rubin, in a teleconference organized for the press by the American Association for the Advancement of Science (AAAS) on Tuesday. Rubin was talking about a paper that appears in this week’s issue of Science, which describes the sequencing and analysis of genomic DNA from the fossilized femur of a middle-aged Neanderthal man who lived about 38,000 years ago.
Rubin was joined on the call by Svante Pääbo of the Max Planck Institute for Evolutionary Biology in Leipzig, Germany, which reported its findings in the British journal Nature. To find their elusive genetic material, the two teams of scientists teased the first “base pairs” — the chemical building blocks of DNA — from the fossilized thighbone of a Neanderthal man who had lived in a cave named Viindija near Vilna in what is now Croatia.
In their Science paper, Rubin and his co-authors concluded that that the genomes of modern humans and Neanderthals are at least 99.5-percent identical, and that the two species last shared a common ancestor approximately 700,000 years ago. Homo sapiens and Homo neanderthalensis split into two separate species about 400,000 years ago (by comparison, humans and chimpanzees diverged about 6 million years ago).
Despite their genetic similarity and cohabitation in the same geographic region for thousands of years, there was no evidence of any significant crossbreeding between the two hominids.
“In this study, we demonstrated that Neanderthal genomic sequences can be recovered using a metagenomic library-based approach, and that specific Neanderthal sequences can be obtained from such libraries,” Rubin said. “We predict that in the near future anthropologists will be able to develop hypotheses about our extinct ancestors through the scanning of billions of base pairs of DNA sequences available on the web rather than only being able to study the limited number of bony remains and associated artifacts that are available in hard-to-access museum collections and field sites.”
The title of the Science paper is “Sequencing and Analysis of Neanderthal Genomic DNA.” Co-authoring with Rubin were James Noonan, Graham Coop, Sridhar Kudaravalli, Doug Smith, Johannes Krause, Joe Alessi, Feng Chen, Darren Platt, Pääbo and Jonathan Pritchard.
In the summer of 1856, a partial skeleton of a hominid was discovered in a cave in the Neander Valley of Germany. Dubbed “Neanderthal” man, the skeleton generated enormous public curiosity. That public interest in our extinct hominid cousin continues to run deep is reflected in the dozens of news stories that have appeared these past two days about this latest Neanderthal DNA sequencing research. Unlike earlier genetic studies of Neanderthal, which involved mitochondrial DNA, the results reported in Science and Nature were based on nuclear DNA, which makes up the vast majority of the genome and contains almost all of the genes.
“Nuclear DNA is where all the biology is,” said co-author Noonan, a post-doctoral fellow in Rubin’s research group who holds joint appointments with Berkeley Lab and JGI. “If you want to understand how traits like language and cognition are encoded, you have to study nuclear DNA.”
Studying ancient DNA from fossilized material represents a major challenge. As a fossil ages, its DNA is degraded by chemical processes. It also becomes contaminated with DNA from the microbes that colonize both the fossil and its immediate environment, and by other organisms, including the humans who handle the fossil.
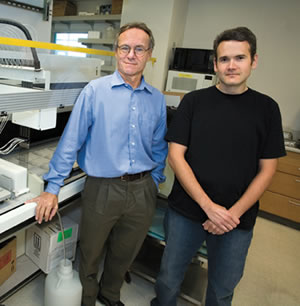

Edward (Eddy) Rubin (left) is a research geneticist and the director of the Joint Genome Institute. James Noonan (right) is a post-doctoral fellow in Dr. Rubin’s research group.
Rubin, Noonan and their colleagues met the fossilized DNA challenge with a unique solution that’s been described as a “targeted approach.” Essentially, they “immortalize” all of the DNA in a fossil sample into metagenomic libraries where individual fragments of the ancient DNA are propagated in microbes. Specific DNA sequences of interest can then can be fished out of this metagenomic library and targeted for study.
“Once the Neanderthal DNA is cloned using our method, we have it forever, and we can recover specific sequences from the library whenever they are needed,” said Noonan.
Through a combination of the sequencing technologies deployed in the Human Genome Project, plus a new massively parallel pyro-sequencing technology, in which enormous amounts of DNA sequence are rapidly and inexpensively generated, Rubin and Noonan and their colleagues were able to recover 65,250 base pairs of Neanderthal DNA from the approximately 6 million base pairs of contaminating DNA in the fossil. A critical factor in helping to confirm that the recovered DNA was Neanderthal rather than human was the short length of the individual Neanderthal sequences — about 50 to 70 base pairs in length, compared to the hundreds to thousands of base pair lengths of contemporary human DNA.
Rubin believes that sequencing the Neanderthal genome should do for paleoanthropology what decoding hieroglyphics did for Egyptology.
“While we learned a lot about the ancient Egyptians from the pyramids and other artifacts, it was only when we were able to decode the squiggles on their artifacts that we began to learn about their everyday life and about their commerce,” Rubin said. “With the sequencing of the Neanderthal genome, the field of paleoanthropology is going to go from data poor to data rich.”
Architect Swims to Maintain Health, and Healthy Attitude
For most of us, getting out of bed, showering, and grabbing a cup of coffee on the way to work is about all we can muster each morning. Kirk Haley, on the other hand, jumps into a chilly pool nearly every day at 6 a.m. and swims for an hour before heading to his job at the Lab.
“It’s hard,” he says of the ritual. “But I feel so much better when I do it.”
 Haley prepares for his work day with early-morning laps at a pool in Moraga.
Haley prepares for his work day with early-morning laps at a pool in Moraga.
Haley is a project manager for the Facilities Division and oversees the funding, design, engineering, and construction of Lab buildings. An architect by training, Haley came to the Lab in 1978 as a junior at Cal to work as a drafter. In addition to Haley’s nearly 30 years at the Lab, his dad also worked here for 40-plus years with the EH&S Division.
Among the projects he’s managed are the construction of the Joint Genome Facility in Walnut Creek and moving the National Energy Research Scientific Computing Center to Oakland. He’s currently working on the design of a guesthouse for visiting researchers and a new computing building, which will house, among other groups, the Oakland Scientific Facility.
With this line of work comes a lot of pressures — unexpected delays, budget issues, and competing deadlines, to name a few. But swimming helps Haley keep a healthy perspective.
“When I’m doing my laps in the early morning hours, there is no e-mail, phone calls, or meetings,” he explains. “I’m able to think about that day’s agenda and sort things out without interruption.”
Swimming has been a part of Haley’s life for a long time. A Lafayette native, he competed on teams throughout his school years. His specialties were the mid-distance freestyle and the butterfly races. He also played water polo.
He skipped water sports in college to focus on academics, but missed it so much, he got back into it shortly after graduating, joining a Masters Swimming team.
Seeking new water adventures, he began participating in open water swimming in the early 1980s. He’s raced in the Bay as well as Lake Tahoe and Donner Lake.
“It can be tough dealing with the elements,” says Haley of the extreme conditions. “Chilly water, big waves, and wind can make it pretty challenging.”
His most recent competition was the Tiburon Mile, which covers the stretch of water between Angel Island and the Tiburon peninsula. And for the first time ever, he competed alongside his 16-year-old son Lincoln, who finished 41st out of 700 swimmers. Dad placed 63rd. Not bad considering some of their fellow competitors were former Olympians.
So what goes through Haley’s mind during these lengthy races?
“At the start, I’m focused on setting a good pace and position, and staying calm. By the middle of the race, I’m wondering why on earth I decided to do this race and how much longer do I have to keep this up. When the end is finally in sight, it seems like it takes so long to get there. But a few days later, after I’ve recovered and shared stories with friends and family, I find myself planning for the next race.”
Modeling a Satellite: SNAP at Full Scale
A space satellite is even more complicated than it looks in drawings, says Robin Lafever of the Engineering Division, “and some problems aren’t easily solved in CAD” — computer-aided design — so the SuperNova/ Acceleration Probe Collaboration is building a one-for-one mockup of the proposed SNAP satellite to make sure everything fits the way it’s supposed to, before the design is finalized. “We’re starting with the lower optics bay of the telescope (1), using the model to work out the plumbing and wiring, thermal strapping, and so on,” Lafever says. “The focal plane (2) is a thorny problem, a real hair-puller.” The full-scale model is being built with Ultralite particle board and Masonite that closely approximates the thickness of the carbon fiber structures that will be used in the real spacecraft.
Engineering’s Matt Hoff (3) attaches mounting hardware to a skeletal representation of the radiation shield that will cover the focal plane (4), identified by its brightly colored stand-in CCDs. Bobby Besuner (5), an engineer from UC Berkeley’s Space Sciences Lab (SSL), joins a bathroom mirror to a particle-board honeycomb to model SNAP’s fold-flat mirror. SSL’s Pat Jelinski, an astrophysicist with long satellite-building experience, helps Lafever and Besuner fix the tertiary mirror in the optical bay (6). Next the team will model the telescope itself, including the primary and secondary mirrors and their mounts, and “sketch in the rest,” including light baffles and the satellite’s outside skin. Lafever hopes industrial partners will supply a model of the basic spacecraft (rockets, etc.) on which the telescope is mounted. Photos by Roy Kaltschmidt, CSO






Berkeley Lab View
Published once a month by the Communications Department for the employees and retirees of Berkeley Lab.
Reid Edwards, Public Affairs Department head
Ron Kolb, Communications Department head
EDITOR
Pamela Patterson, 486-4045, pjpatterson@lbl.gov
Associate editor
Lyn Hunter, 486-4698, lhunter@lbl.gov
STAFF WRITERS
Dan Krotz, 486-4019
Paul Preuss, 486-6249
Lynn Yarris, 486-5375
CONTRIBUTING WRITERS
Jon Bashor, 486-5849
Allan Chen, 486-4210
David Gilbert, (925) 296-5643
DESIGN
Caitlin Youngquist, 486-4020
Creative Services Office
PHOTOGRAPHY
Roy Kaltschmidt, 486-5731
Creative Services Office
Berkeley Lab
Communications Department
MS 65, One Cyclotron Road, Berkeley CA 94720
(510) 486-5771
Fax: (510) 486-6641
Berkeley Lab is managed by the University of California for the U.S. Department of Energy.
Online Version
The full text and photographs of each edition of The View, as well as the Currents archive going back to 1994, are published online on the Berkeley Lab website under “Publications” in the A-Z Index. The site allows users to do searches of past articles.
Flea Market is now online at www.lbl.gov/fleamarket
Molecular Foundry — ‘Living Laboratory’ for Nano Research — Earns Energy Efficiency Honors
Long before Berkeley Lab’s newest monument to cutting-edge research — the Molecular Foundry, a state-of-the-art nanotechnology building — began construction, planners and architects vowed to make this a showcase facility for energy efficiency.
Now, open for just a few months, the Foundry has already earned its first recognition for cost-effective energy technologies and practices. The Federal Energy Management Program has given one of its Federal Energy Saver Showcase Awards to the Department of Energy’s newest nanoscience center. Berkeley Lab was the only national research lab to receive one.
The bottom line is this: the innovative features in the Foundry that minimize energy use have resulted in an estimated annual savings of about 8,500 million Btu, or a cost savings of about $55,000. Not to mention the environmental benefits that result.
“We all have to do something about carbon emissions, and energy is the dominant part of that,” said Foundry Project Manager Joe Harkins. “Every little bit helps, and laboratory buildings are very energy-intensive.”
This one was going to be a special challenge. The $85 million, six-floor, 90,000-square-foot hillside structure would house extremely sensitive instrumentation requiring significant heating and cooling, a 5,000-square-foot clean room with high-filtered air requirements, and lots and lots of office and laboratory space.
Once Project Director Jim Krupnick set the goal of achieving a “LEED Silver” designation, Laboratory project managers and facilities experts, the San Francisco architectural firm of SmithGroup, and an energy consultant from Gayner Engineers set out to find the magic balance between energy conservation and scientific requirements. LEED certification is the recognized standard for measuring building sustainability. The LEED green building rating system — developed and administered by the U.S. Green Building Council in Washington D.C. — is intended to promote design and construction practices that increase profitability while reducing the negative environmental impacts of buildings and improving occupant health and well-being.
The Molecular Foundry is the first building at Berkeley Lab designed to meet strict LEED criteria.
Innovations can be found from basement to roof, but start with the fume hoods.
A fume hood is a large cupboard-like enclosure designed to limit a scientist’s exposure to hazardous fumes. A fan circulation system draws air from the laboratory through the fume hood to prevent build-up of hazardous fumes. One of these units generally uses as much energy at three houses. And the Foundry has 42 of them.
“We spent a lot of time studying the sash design of the fume hood,” said Steve Greenberg, an engineer in Environmental Energy Technologies who consulted on the project design. “Horizontal safety shields were installed to block the openings of the hoods and thus saving on the amount of air you are exhausting. That reduced the available space, and thus the total air flow, by 20 percent.”
The scientists who were going to work in these variable-volume fume hoods were given a demonstration at UC San Francisco’s Genentech Hall across the bay, and they gave the concept a thumbs-up.
The clean room was a different challenge. Very energy-intensive, these rooms traditionally have high air-flow rates, a sophisticated filtration system, and humidity controls. A “class 100” clean room like the one in the Foundry can’t have more than 100 particles of 3-micron size within a cubic foot of air, so that sensitive and precise photolithography can be performed without contamination.
So the planning group borrowed concepts used in EETD’s high-tech buildings program and custom-designed the 150 fan filter units with the highest efficiencies, and with modularity that allowed flexible replacement, repair and use. In addition, double-paned exterior windows were coated with a film that blocks ultraviolet rays, allowing daylighting and views — a unique feature for a clean room.
For water conservation and efficiency, a continuous cooling water loop eliminated the traditional once-through approach, and a vacuum system reduced the need for energy-consuming aspirators. Outside, landscaping is dominated by succulents and native grasses requiring minimal watering.
But the most talked-about innovation is the waterless urinal, 12 of them installed in the men’s bathrooms. Each configured with a replaceable odor trap, the urinals eliminate the one-gallon-per-flush requirement of common low-flow urinals.
Lab planners used the concept of “right-sizing” when determining utility load requirements. That is, as Harkins explained it, “We measured electrical levels in buildings with similar uses, like 62 and 66, and used that base for our assumptions in the Foundry.” Thus mechanical and electrical equipment could be sized intelligently. As a result, the size of the electrical system was reduced by 38 percent and the HVAC system by about a third, reducing “first costs” by $2.5 million. Air handlers and chillers were reduced; boilers and cooling towers were downsized. The Foundry’s energy performance beats the state and federal standards by 25 to 35 percent.
Green concepts were used during construction, too. About 80 percent of the waste generated was recyclable. Care was taken to install rapidly renewable materials, like the wood-bamboo flooring. All the lights have occupancy sensors and high-efficiency ballasts. Almost a mile of ductwork was precisely cleared, cleaned and capped. The carpets and paint were carefully chosen so that they would not “out-gas” (i.e., give off vapors or volatile organic compounds). Refrigerants in the chillers have zero ozone-depletion potential.
A case study by the Environ-mental Protection Agency and the DOE’s FEMP program declared that the Foundry “can serve as a ‘living laboratory’ for developing and refining energy-savings strategies for high-technology buildings.”
Besides Harkins and Greenberg, the e-team included Project Director Krupnick, who with Harkins accepted the FEMP award in Washington last month; Facilities’ Doug Lockhart; EETD’s Dale Sartor and his applications group; consulting engineer Nick Mironov of Gayner Engineers; SmithGroup’s Irene Monis; and Martyn Dodd of Energy-Soft.
Berkeley Lab has applied for the LEED “Silver” designation and expects to hear the results in December.



Molecular Foundry researcher Dmitri Talapin (top) works inside one of the variable-volume fume hoods designed specifically for the facility to reduce air flow and save energy. The Foundry's clean room (center) features custom-made fan filter units and natural daylighting. Floors like this one (left) are made of renewable wood and bamboo materials.
Lab Directed Research and Development Awards Totaling More Than $16 Million are Announced
Berkeley Lab Director Steven Chu announced the awards for the FY2007 Laboratory Directed Research and Development (LDRD) program (see the View for Dec. 16, 2005). A total of over $16 million in operating and capital equipment (including G&A) is for 78 projects from a field of 159 proposals. Of these, 31 are new and 47 are continuation projects. The evaluations used input from divisional reviews as well as laboratory review committees.
In making the awards, Director Chu said, “Scientists for all proposals are again commended for their creativity in offering many promising directions for Berkeley Lab. We were very pleased at the high quality of all the proposals.”
| Lead-PI | Title | Allocation ($K) |
Adams |
Research Tools for the Conversion of Cellulose to Ethanol: Structural Studies of Cellulose Synthesis | 150 |
Andersen |
Microarray Technology for Fungal Identification: Its Utility from the Environment to Homeland Security, Biomedicine, and Beyond | 147 |
Arenholz |
Development of Fast Switching Superconducting Magnets for Spectroscopy Applications | 251 |
Arnold |
Synthetic and Electrochemical Approaches to Metal-Metal Bonds in Actinides | 69 |
Battaglia |
Advanced Monolithic Silicon Pixel Detectors | 272 |
Bauer |
Soft-collinear Effective Theories applied to Collider Physics | 285 |
Bell |
Structured, Adaptive Mesh Refinement Method for Multiphase Reactive Transport in Groundwater | 219 |
Bergman |
Conversion of Glycerol and Aromatic Compounds from Biomass to Major 3- and 6- Carbon Industrial Organic Compounds | 127 |
Berket et al |
On-demand Overlays for Scientific Applications | 129 |
Berryman |
Applications of Adjoint Field Methods and Time-Reversal Data Processing to Inverse Problems in Electromagnetics, Seismics, and Ultrasonics | 147 |
Bluhm, Wilson |
Chemical Reactions at Liquid/Vapor Interfaces Probed by Photoemission Spectroscopy | 146 |
Butland |
Functional Interatomics: Integrating Physical and Functional Interaction Networks | 350 |
Cate, Marletta et al |
Cellulose Degradation and Cellulosomes | 339 |
Chevassut |
Cryptographic Foundations for New-Generation Distributed Systems | 73 |
Clark |
New Experimental Initiative to Deduce (n,f) Cross Sections for Advanced Fuel Studies | 147 |
Coates et al |
An Investigation of the Microbial Processes Involved in Electron Transfer Onto the Anode of a Biological Fuel Cell | 276 |
Cooper |
Transcription Cofactor PC4 Interactions with RNA Polymerase and XPG in Transcription-Coupled Repair | 288 |
Crooks |
Statistical Dynamics of Protein Evolution | 143 |
Denes et al |
Novel Imaging Detectors for Materials and Biology | 529 |
DePaolo |
Microcharacterization and Chemical Microdynamics of Atmospheric Mineral Dust | 136 |
Destaillats et al |
Understanding the Chemistry of Innovative Air Cleaning Technologies | 108 |
Eisen |
Computational and Experimental Testing of Methods for Binning Sequences from Metagenomic Studies | 160 |
Fawley et al |
FEL Concepts for Multiple Independent X-ray Beamlines | 292 |
Francis et al |
Tailoring the Self Assembly of Functionalized Biomolecular Building Blocks | 202 |
Frei et al |
Biological and Nonbiological Integrated Systems for Solar to Fuel | 366 |
Gadgil |
Arsenic Electrochemistry | 132 |
Gilbert |
Behavior and Impact of Nanoparticles in the Environment | 160 |
Guo |
Building In-Situ Electronic Structure Study Capability with Photon-in/ Photon-out Soft X-ray Spectroscopy |
252 |
Holman |
Compositional and Functional Analysis of Cell Walls During Metal-Bacterial Actions | 174 |
Hugenholtz |
Metagenomics-Enabled Analysis of Termite Hindgut Microbiota for Biomass Conversion and Cleaner Energy | 115 |
Javey |
Integration of Synthetic Nanomaterials for High Performance, Robust, and Flexible Circuitry | 147 |
Ji |
Ultra-compact Field Desorption Neutron Source for Cancer Research | 204 |
Kaindl |
Terahertz-frequency Conductivity and Ultrafast Optical Excitations in Single-Walled Carbon Nanotubes | 95 |
Kirchstetter |
Soot in Ice: Does Soot Enhance the Melting of Ice? | 163 |
Konerding |
Software Application Infrastructure for Efficiently Managing Large-Scale Computational Biology Experiments | 136 |
| Kostecki, Mao | Surface Plasmon-Enhanced Photovoltaic Device | 108 |
Lee (S.-W.) |
Fabrication of Photovoltaic Devices Using Nanostructured Biomaterials | 191 |
| Lesko, Wang | Physics Detector and Sensor Technologies Applied to Geological and Geophysical Applications at DUSEL | 438 |
| Lester et al | Development and Application of Quantum Monte Carlo (QMC) Methods to Biological Systems | 218 |
Logan et al |
Enabling High Energy Density Physics at LBNL | 424 |
Marcus et al |
Measurement of Molecular Shape and Assembly Using X-ray Scattering | 494 |
Markowitz et al |
Integrated Microbial Community Genomes Data Management System | 436 |
Martin et al |
Left-handed Nanoscale Meta-materials: Towards the Optical Domain | 206 |
McMahon et al |
Integrated Decision Support Tool for Joint Optimal Control of Energy and Water Systems Under Uncertainty | 216 |
Melis, Mehlhorn |
Bio-oil Accumulation in Unicellular Green Algae: A Pilot Project | 140 |
Moridis |
Interrelation of Global Warming and Hydrate Dissociation in Oceanic Accumulations | 147 |
Murayama |
New Directions for Theoretical Physics at the TeV Scale | 362 |
Nakagawa |
New Technology for Permeability Enhancement for Natural Gas Extraction in Tight Reservoirs | 116 |
Newman et al |
Photons to Fuels - the Electrochemical Reduction of Carbon Dioxide to Methanol | 357 |
Oldenburg |
Coupled Modeling of Hydrology, Nutrient Cycling, and Vegetation: Applications to Water Quality and Water Balance | 125 |
Oliker |
Enhancing Commodity Scalar Processors with Vector Components for Increased Scientific Productivity | 260 |
Pennacchio |
Development of Cost Effective Sequence-based Technologies to Identify Genomic Alterations in Cancer | 212 |
Phair |
Improved Spectroscopy of Weakly Bound States in Nuclei | 301 |
Pinar et al |
Advanced Computational Tools for Electric Power Systems | 161 |
Prausnitz |
Properties of New Ionic Liquids for Electrochemical Applications and for Extraction of Heavy-metal Cations from Wastewaters | 80 |
Ritchie |
Aging, Disease and the Mechanical Response of Biological Tissues, Specifically in Human Bone | 146 |
Romano |
Statistical Feature Modeling for Scientific Data Via Basis Decomposition | 116 |
Rotenberg |
NanoARPES: A New Detector for Nanometer-scale Electronic Structure Measurements | 289 |
Schaffer et al |
Cooperation of Biochemical and Mechanical Signals in Regulating Cell Fate Decision During Tissue Morphogenesis | 246 |
Schild |
Determine if PIR51 is a Potential Tumor Suppressor Gene Similar to BRCA2 | 115 |
Schlegel |
Baryon Oscillations and Dark Energy: Prototyping Instruments | 140 |
Schroeder |
Laser-plasma Accelerator Driven Free-Electron Laser with High-harmonic Seeding | 144 |
Shalf |
Power Efficiency Metrics for High Performance Computing | 141 |
Sichtermann |
Hyperons in Polarized Proton Collisions and the Origin of the Nucleon Spin | 147 |
Skinner |
Integrated Performance Monitoring of Grid and HPC Workloads | 173 |
Somorjai |
Electron Flow Generated by Gas Phase Exothermic Catalytic Reactions Using Metal-Semiconductor Nanodiodes | 146 |
Stamper-Kurn |
Studies of Quantum Antiferromagnetism in Two-dimensional Triangular Lattices Using Ultracold Atoms | 121 |
Steefel |
Biochemical Reaction Rates and Pathways in Porous Media | 88 |
Tilley |
New Approach for the Catalytic Conversion of Methane and Other Inert Hydrocarbons | 112 |
Tyliszczak |
Versatile Mini-scanning Transmission X-ray Microscope (mSTXM) | 151 |
Vetter |
Conceptual Study for a Novel Nuclear Astrophysics Accelerator Capability | 144 |
Wan et al |
High Brightness Cathodes as Electron Sources for FELs | 286 |
Wilkening |
Extended First Order System Least Squares Finite Elements | 54 |
Wyrobek |
Expression Profiling of Radiation and Cancer Susceptibility Genes | 399 |
Yang et al |
Hierarchically Nanostructured Systems for Solar Energy Hydrogen Production | 329 |
Yashchuk |
Ultra-high Resolution Optics for Soft X-ray Inelastic Scattering | 141 |
Zholents et al |
Emittance Manipulation and Beam Conditioning for FELs | 288 |
Zolotorev |
Low Energy-spread Electron Source | 182 |
Total Allocations $16,019 |
Legends and Heroes: Lab and Guests Pay Tribute to Its Legacy
Historians of the 23rd century will call it “the age of heroes,” retired UC Berkeley scientist John Heilbron said, an age that had its Achilles (Ernest Lawrence), its Ulysses (Luis Alvarez) and its Jason (Glenn Seaborg). These pioneers and their colleagues, he said, populated “the Olympus of Nuclear Science,” a place that in 75 years has achieved lofty stature in a variety of fields.
Heilbron was just one of many who sang the praises of Lawrence Berkeley National Laboratory on the occasion of its 75th anniversary earlier this week, both in a daylong scientific symposium and at an evening gala dinner. Energy Secretary Samuel Bodman, in a dinner address, called the Lab “truly sacred ground for American science and engineering.” The other testimonials and reminiscences supported that label.
The View was just going to press as events of Nov. 14 concluded, so a complete report of the anniversary celebration will appear in the December issue. In the meantime, some of the drama was captured in these images by photographer Roy Kaltschmidt.
 |
 |
|
Lab Director Steve Chu and Nobelist George Smoot hold street signs presented to them by Deputy Director Graham Fleming (center), in honor of the roads that were named after them on the Hill. |
(From left) Former Berkeley Lab Directors Andy Sessler and Chuck Shank join current Director Steve Chu in discussing Lab past, present and future at the symposium. | |
 |
 |
|
UC Berkeley Chancellor Bob Birgeneau shares a toast to the Lab’s 75th with about 250 others at the Claremont Hotel dinner |
Former Deputy Director Pier Oddone returns to share the “enduring values” of national labs, established at Berkeley | |
 |
 |
|
| Historian John Heilbron speaks of heroes and Elysian Fields | Energy Secretary Samuel Bodman speaks at gala dinner | |
 |
 |
|
| University of California President Robert Dynes: Lab success “uniquely Californian” | Distinguished Scientist Mina Bissell explains “the architecture of life” |
Berkeley Lab Science Roundup
Living High on Sugar
 Whole genome comparisons of bacteria that produce lactic acid reveal how they add zest to many foods and could help with biofuels and other special chemicals.
Whole genome comparisons of bacteria that produce lactic acid reveal how they add zest to many foods and could help with biofuels and other special chemicals.
Bugs made the news last month, some as familiar as the bacteria that ferment wine, cheese, sourdough bread, and yogurt, others as exotic as extremophiles living two miles underground in the hot darkness of a South African gold mine. Sequences of nine species of lactic acid-producing bacteria were compared by the Genomics Division’s Paul Richardson, head of JGI’s Genomics Technologies Program, and his colleagues in a group led by David Mills of UC Davis. Richardson says the study, published in the Oct. 17 issue of Proceedings of the National Academy of Sciences, “offers a window into the sugar metabolism and energy conversion systems” of these high-living critters, which could lead to improved methods of food production.
Living Low in the Rocks
The bugs in the South African mine, by contrast, subsist on sulfates and hydrogen split from salty water by radioactivity in the surrounding rocks. Eoin Brodie, Gary Andersen, Terry Hazen, and Todd DeSantis of Earth Sciences were members of the multi-institutional team reporting in the Oct. 20 Science. Hazen says the denizens of a formation “probably tens of millions of years old” are Firmicutes bacteria that can’t be grown in the laboratory but which “have the ability to adapt to anything that comes their way.” If there’s life on Mars, it may be a lot like Firmicutes.
Chemicals by Contrast
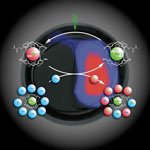
A high-contrast difference image is obtained by subtracting the signal of hyperpolarized xenon caged in targeted biosensors (green) from the signal indicating polarization transfer to free xenon (blue to red).
Also described in the Oct. 20 Science is a revolutionary new technique for magnetic resonance imaging. Molecular MRI is invaluable in medical diagnosis and biological research, but traditional MRI suffers from low sensitivity and slowness. Alexander Pines of the Materials Sciences Division and David Wemmer of the Physical Biosciences Division, with Leif Schröder, Thomas Lowery, and Christian Hilty, have demonstrated molecular MRI that is 10,000 times more sensitive, 3,000 times faster, and capable of detecting more than one kind of molecule simultaneously. Their demonstration of the technique, dubbed HYPER-CEST, combines hyperpolarized xenon atoms caged in tailor-made biosensors with chemical exchange saturation transfer (CEST) to enhance image contrast.
Getting to the Baryon Bottoms
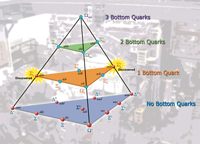
The CDF detector at Fermilab has found two high-spin baryons containing bottom quarks. Other such baryons are predicted (blue circles) but have yet to be detected.
Lina Galtieri and Marjorie Shapiro are among the many Berkeley Lab physicists who are members of the 61-institution, 700-person CDF collaboration at Fermilab, which on October 23 announced the discovery of two new baryons predicted but never seen before. Called sigma-sub-b’s, the massive particles — one positively and one negatively charged — are relatives of ordinary protons and neutrons, and like them consist of three quarks. But the new baryons are six times heavier, contain a bottom quark as well as up or down quarks, and survive for only a fraction of a second in the debris of Tevatron collisions. CDF researchers expect to find more kinds of such heavy baryons.
Slippin’ and Slidin’ at the ALS
A team led by John Tainer of the Life Sciences Division and the Scripps Research Institute, including Berkeley Lab’s Greg Hura and Scott Classen, used the SIBYLS beamline at the ALS to elucidate a complex molecular machine, part ligase, part “sliding clamp,” which can replicate, recombine, and repair DNA by wrapping itself around the double helix and recruiting other proteins. Tainer says the results, published in the Oct. 20 issue of Molecular Cell, show for the first time “how these proteins can dynamically assemble and change their shape to join DNA ends during replication and repair.”
Curling the Mesenchymes
Weightlifters exercise differently than long-distance runners, and so do mesenchymal stem cells, destined to become different kinds of tissue like bone or muscles. The Life Sciences Division’s Song Li and colleagues grew a layer of stem cells inside a mechanism that could stretch them, with periodicity like the pulse rate of the human body. The stretching increased expression of genes related to the kind of strength needed for the walls of blood vessels or smooth muscle, but decreased expression of genes needed to withstand compression, as in bone. Findings were published online Oct. 23 in Proceedings of the National Academy of Sciences.
Switching on the Light
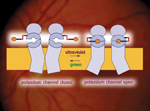
A molecule that changes shape when illuminated by light of different colors can act as a switch to open or close potassium channels and activate nerve cells like those in the retina.
Richard Kramer and Dirk Trauner of the Materials Sciences Division and Ehud Isacoff of Physical Biosciences, with colleagues at UC Berkeley, Stanford, Caltech, and the Scripps Institution of Oceanography, are developing molecular tools they hope can cure blindness, activate drugs on command, and stimulate nerve cells at the flash of a colored light. The team has altered ion channels in nerve cells to open or close in response to green or ultraviolet light, described earlier this year in Nature Chemical Biology. As members of the new Berkeley Lab and UC Berkeley Nanomedicine Development Center sponsored by NIH, announced Oct. 31, their goal is to equip victims of degenerative blindness with light-sensitive photoswitches in the retina.
Stickiness and High I.Q.
Human brain cells are stickier than those of our cousins the apes and monkeys, which may be why the human brain is capable of more complex cognitive functions. By comparing the entire genome of many organisms to that of humans “we were able to identify a series of human-specific sequence changes that have a high likelihood of turning genes on and off,” says Eddy Rubin, director of Berkeley Lab’s Genomics Division and DOE’s JGI. The changes all point toward greater neuronal-cell adhesion. Rubin, James Noonan, and Shyam Prabhakar report their findings in the Nov. 3 issue of Science.
Oxygen From Sunlight and Water
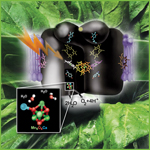
The arrangement of manganese, oxygen, and calcium atoms in the manganese complex is key to how Photosystem II can split water into oxygen atoms and hydrogen ions during photosynthesis.
The Nov. 3 Science also reports the most detailed structure yet of the manganese complex, a cluster of manganese, oxygen, and calcium atoms in the protein machinery called Photosystem II. Billions of years ago primitive bacteria began using solar energy to split water into hydrogen and oxygen during photosynthesis; it’s why plants and some microorganisms exhale oxygen and make life possible for the rest of us today. Vittal Yachandra and Junko Yano of Physical Biosciences led an international collaboration of scientists including Berkeley Lab’s Kenneth Sauer and Yulia Pushkar, using high-resolution synchrotron studies of the manganese complex to show just how the water-splitting is done.
Physicist Michael Ronan Remembered for His Hard Work, Versatility, and Enthusiasm
Physicist Michael Thomas Ronan, a versatile experimentalist who made major contributions in detector design and data analysis, fell to his death on the afternoon of Tuesday, October 17, from an upper floor of Bldg 50B. He was 57.

Ronan joined Berkeley Lab’s Physics Division in 1976, shortly after receiving his Ph.D. in elementary particle physics from Northeastern University in Boston. He joined a team headed by Lina Galtieri, hunting for charmed particles at SLAC’s SPEAR ring.
“Mike would go out of his way to do the best job on the electronics; later he did a fantastic job analyzing the data,” says Galtieri, noting his important contributions to an experimental run that culminated in the discovery of a new resonance, the psi(3772).
Ronan then helped build the first Time Projection Chamber (TPC), devised by David Nygren, and was in charge of the trigger and timing system.
Nygren says, “He created a sliding time window in the circuitry that was quite sophisticated for the times. I marveled at his ability to move seamlessly back and forth between hardware and software.”
Ronan became a champion of the TPC concept, and in recent years spearheaded a TPC detector design for the International Linear Collider, serving as the Americas representative for the Global Large Detector collaboration.
“Perhaps someday we will be recording collisions in a time projection chamber, just as Mike had envisioned,” says ILC Global Design Effort director Barry Barish. “Mike was a colleague and a brilliant experimentalist, who had made a deep commitment to the ILC.”
Colleague Willy Chinowsky met Ronan on the Lab shuttle bus some weeks before his death and says, “he seemed happy and pleased with progress of the introductory physics class he was teaching on campus.”
Later, however, Ronan told his Physics 8A students he was suffering from depression and would not continue to teach the class, according to the Daily Californian. Yet on the afternoon of his death, friends say he was making plans for the evening. The exact circumstances of his fall are still under investigation by the University of California Police Department.
Ronan is survived by his wife, Alda, and his grown children Kevin and Stacey.
Alpen, Radiation Biophysicist, and Former ALD, Donner Lab Director, Dies
Edward Alpen, a distinguished radiation biophysicist with Berkeley Lab, passed away Nov. 3 at the age of 84. He had suffered from complications as a result of radiation therapy for a brain tumor.

Alpen joined the Lab in 1975 as director of the Donner Laboratory and associate director of the Biology and Medical Division. In 1986, he transitioned to a senior faculty scientist of cell and molecular biology, while holding professorships at UC Berkeley and UC San Francisco. He retired from the Lab in 1991, though was temporarily reappointed to continue PI work on two research programs, which ended in 1995.
His principal areas of research were experimental radiotherapy with charged particle beams and neutrons, radiation biophysics and medical physics, radiation, carcinogenesis, non-stochastic late effects in organ systems, and cellular radiation biology.
He received both his B.S. and Ph.D. from UC Berkeley.
The recipient of many awards, including the Distinguished Service Medal of the Department of Defense and the Navy Science Medal for Distinguished Achievement in Science, Alpen was also the author of the highly praised book Radiation Biophysics.
People, Awards, and Honors

A University IT Expert To Lead Lab Division
Rosio Alvarez, a California native with a distinguished record as a university technology manager and expert on organizational impacts of technology, has been named Berkeley Lab’s Director of the Information Technology Division and Chief Information Officer. Effective Jan. 1, she will succeed current IT Director Sandy Merola, who was recently named Deputy Chief Operations Officer. Alvarez was introduced Tuesday to IT division staff by Lab Director Steve Chu, who cited her extensive publication and presentation portfolio and her regional IT leadership in the northeast U.S.

New HR Head Brings Public, Private Expertise
Vera Potapenko, Berkeley Lab’s new Chief Human Resources Officer and Department Head for Human Resources, brings almost 30 years of experience in both public and private sectors, and a Ph.D. in organizational communication and rhetoric, to her role. In announcing the appointment, Chief Operations Officer David McGraw cited her skills and accomplishments “in human resource management and strategic planning, organizational design and effectiveness, and leadership development.” Potapenko started this week after leaving Northwestern University as its Director of Human Resources and Staffing.

Library Adds New Reference Librarian
Michael Golden is the Lab’s new reference desk librarian. Golden has a Masters degree in Library and Information Science and eight years experience as an academic reference librarian at University of Missouri, Kansas City, as well as other campuses. He has an additional five years experience as a bench scientist in chemistry.

Former Staff Activities Coordinator Back at Post
Arabella Schmidt returns as the new coordinator of the Employee Activities Association (EAA) for FY07. A four-year veteran of the Lab’s Human Resources Office, Schmidt is currently a recruiter for both the Operations and Accelerator and Fusion Research Divisions. “I am looking forward to coordinating this year’s EAA. It’s very important to make sure employees have various opportunities to get to know other lab employees and promote work-life balance,” she said. The EAA sponsors recreational, cultural, educational and social activities at the Lab for employees.

Mathematician is Chair Of Computation Group
John Bell, head of the Computational Research Division’s Center for Computational Sciences and Engineering, has been elected chair of the Computational Science and Engineering activity group for the Society for Industrial and Applied Mathematics (SIAM). According to the SIAM website, the activity group "fosters collaboration and interaction among applied mathematicians, computer scientists, domain scientists and engineers in those areas of research related to the theory, development, and use of computational technologies for the solution of important problems in science and engineering."

Materials Scientist Wins APS’s Langmuir Award
Berkeley Lab materials scientist Gabor Somorjai has been awarded the 2007 Irving Langmuir Prize in Chemical Physics from the American Physical Society. The prize was established to recognize and encourage outstanding interdisciplinary research in chemistry and physics, in the spirit of 1932 Nobel Laureate Irving Langmuir. According to the APS, Somorjai was awarded the prize “for his pioneering research in surface chemistry and delineation of catalytic mechanisms.”

 L-R: Walukiewicz; Man Yu
L-R: Walukiewicz; Man Yu
Researchers Get ‘Most Promising’ Tech Award
Wladek Walukiewicz and Kin Man Yu of the Materials Sciences Division got a surprise at the annual R&D 100 awards banquet in Chicago, when the editors of R&D Magazine presented them with the “Most Promising New Technology” award for their high efficiency multiband semiconductor material for solar cells. Most semiconductors have narrow band gaps and respond to only certain frequencies, wasting a lot of sunlight. The new materials’ band gaps can be split to respond to a range of frequencies, making use of almost the entire solar spectrum.

Materials Expert Among ‘Scientific American 50’
Antoni Tomsia of the Lab’s Materials Sciences Division was recently included in the “2006 Scientific American 50,” the magazine’s annual list that recognizes people and organizations with accomplishments in research, business, or policymaking that demonstrate outstanding technological leadership. Tomsia was chosen for his development of a multilayered nacrelike material, which could eventually be used in a myriad of applications for which strength and lightness are imperative, such as dental implants.



 L-R: Karsten Pruess; Jonny Rutqvist; Chinfu Tsang; Yvonne Tsang
L-R: Karsten Pruess; Jonny Rutqvist; Chinfu Tsang; Yvonne Tsang
Earth Scientists Lauded For Geological Research
Four Berkeley Lab earth scientists have recently received awards for their research. Karsten Pruess was given the O. E. Meinzer Award from the Geological Society of America. The award recognizes his work in multiphase and multicomponent flow and transport in complex geological media. Jonny Rutqvist, with co-authors Chin-Fu Tsang and Yvonne Tsang, have been honored by the American Rock Mechanics Association with a Case History Award for his analysis of coupled thermo- and hydro-mechanical processes within the Yucca Mountain Drift Scale Heater Test. The test is the largest-scale and highest-temperature field experiment to date, with the objective of studying physical and chemical processes associated with a potential nuclear repository.