|
|
|
| |
 |
|
|
| |
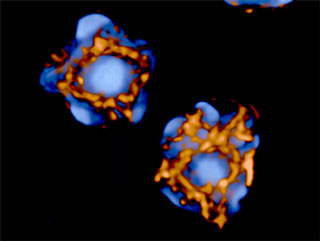
Intricately wound, folded,
and looped chromatin (blue) meets chromatin-remodeling and modifying
factors at sites on a cage-like structure formed by SATB1 proteins
(gold). In this image the chromatin is densely packed heterochromatin,
a type associated with silent genes. (Image: Abby Dernburg) |
|
|
A mammalian body contains trillions of cells, most of them packed with
a whole genome's worth of DNA. Stretched out straight, the DNA in the
nucleus of just one cell would be a yard or two long. How does it all
fit?
Through tight, intricate, twisting and folding: a thread of DNA winds
around a spool made of proteins called histones; thread and spool together
make a nucleosome. The DNA strings the nucleosomes together like beads,
and the beads clump together in thick fibers; the fibers fold into loops,
and the loops are further looped into the ropy mass of chromatin of which
the individual chromosomes in the nucleus are made.
So many levels of winding, folding, and looping create a dilemma: for
a cell to express proteins, it needs to transcribe genes, which requires
double-stranded DNA to unzip where the gene is encoded. DNA wound up tight
in chromatin can't unzip; like the wire in a coiled steel cable, most
of it can't even be reached.
Researchers led by Terumi Kohwi-Shigematsu of Berkeley Lab's Life Sciences
Division are learning the secrets of how specific sites of DNA in the
genome can be made accessible for protein factors that change the chromatin
structure locally. These changes make gene transcription possible or repress
it; in this way, at appropriate times and places, specific sets of genes
are expressed or remain silent, and each type of cell expresses only the
genes appropriate to its physiological role.
Investigating unusual DNA structures
A decade ago Kohwi-Shigematsu and her husband, Yoshinori Kohwi, also
in Berkeley Lab's Life Sciences Division, were investigating certain DNA
sequences with a strong tendency to adopt noncanonical structures — ones
inclined to coil not quite "by the book."
They identified a special class of sequences with a strong tendency to
pop open — and also to unzip the neighboring sequences, when the DNA
helix is under negative supercoiling — that is, when the intact double
strand of DNA is coiled in the opposite direction from the way the two
strands coil around each other. They called these sequences "base
unpairing regions," or BURs.
BURs under negative supercoiling tend to close up and become double stranded
if the microenvironment gets saltier. But short core sequences, a few
bases long, refuse to pair up no matter how salty the surroundings.
BURs are rich in the bases adenine and thymine (A and T), which pair
only with each other (as do the other two DNA bases, cytosine and guanine,
C and G). While sequences rich in A and T separate a bit more easily into
single strands than C- and G-rich sequences, not just any stretch of As
and Ts readily unzips.
Base unpairing regions, however, contain clusters of ATC sequences where
only well-mixed As, Ts, and Cs occur on one strand. Kohwi and Kohwi-Shigematsu
called such a cluster an ATC sequence context.
"We reasoned that if these regions were biologically important,
there must be an important protein associated with them," says Kohwi-Shigematsu.
Using cloned BURs as bait, they went fishing in a library of proteins
and hooked a big one, which they straightforwardly named "special
AT-rich binding protein 1," better known as SATB1.
Although SATB1 is very particular about latching onto base unpairing
regions, it does not attach itself to exposed DNA bases; instead, it slides
into the minor groove on the outside of double-stranded BUR sequences.
Rather than recognizing a particular primary sequence, SATB1 recognizes
the ATC sequence context, a likely site for base unpairing. Thus SATB1
manages to be both specific and versatile at the same time.
|
 |
|
|
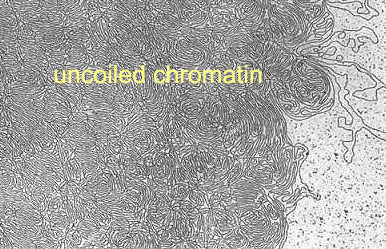
In a strong salt solution the cell nucleus bursts and
chromatin spills out. But even in very strong solutions not all
proteins are removed.
|
|
BURs are often found in matrix attachment regions, operationally defined
as genomic DNA sequences tethered to the nuclear components that resist
salt extraction.
Arming the immune system
Matrix attachment regions in general bind to several proteins, most found
in many different cell types. SATB1 works only in a few distinct kinds
of cells (including the embryonic stem cells much in the news), all of
which are unspecialized precursors of mature cells that later assume particular
functions. SATB1 is most widespread in the cells known as thymocytes.
Thymocytes, so named because they grow to maturity in the thymus gland,
are the precursors of T cells, among the immune system's most potent weapons.
"Killer" T cells (cytotoxic lymphocytes) go straight for the
metaphorical jugular of invading disease organisms, tumors, or other cells
marked for destruction. "Helper" T cells emit proteins like
interleukin 2 that help identify targets, stimulate the defenders, and
aid in the attack. (Helper T cells are themselves a principal target of
HIV infection.)
Mature killer and helper T cells are distinguished by cell-surface markers
designated CD8 and CD4. Early in their development, thymocytes have neither
of these markers. They proliferate rapidly and differentiate into a double-positive
stage, expressing both CD4 and CD8.
During the double-positive stage, cells that are useless or "self-reactive"
— having an unfortunate tendency to kill the host — are eliminated in
droves; approximately 98 percent of the thymocytes generated each day
die without leaving the thymus. Survivors become "single positive"
for either CD4, as mature helper T cells, or CD8, as mature killer T cells.
Kohwi-Shigematsu and her colleagues soon learned that SATB1 plays a crucial
role in T-cell development.
Part 2, A
structural protein that regulates genes
|