
Chemists Look to ‘Hedgehog’ for Brain Damage Therapy
Conference on Women in Science Coming to Lab
Chemists Look to ‘Hedgehog’ for Brain Damage Therapy

Chemical engineer David Schaffer has identified the Sonic hedgehog protein as a “cell fate switch” that can activate the proliferation of adult neural stem cells.
Sonic the hedgehog was such a popular video game hero in the early 1990s that when a new protein whose mutation created spiny-backs in flies was discovered in 1993, it was named after him. The Sonic hedgehog video game may have gone the way of the eight-track cassette or the floppy disk, but the Sonic hedgehog protein has only just begun its rise to fame.
A team of scientists led by chemical engineer David Schaffer has identified the Sonic hedgehog protein as a critical “cell fate switch” that can cause the normally small number of adult neural stem cells in the hippocampus of the brain to proliferate. This discovery holds important therapeutic implications for future treatments of brain damage suffered as a result of Alzheimer’s, Parkinson’s and Huntington’s diseases, as well as strokes or head traumas.
“We have shown that the Sonic hedgehog network of proteins regulates the proliferation of adult hippocampal neural stem cells both in vitro and in vivo,” says Schaffer. “In studies with rats, increasing the presence of Sonic hedgehog proteins in the hippocampus resulted in a 3.3-fold increase in neural stem cell divisions.”
Schaffer holds appointments with the new Synthetic Biology Depart-ment in Berkeley Lab’s Physical Biosciences Division, UC Berkeley’s Chemical Engineering Department, and the Helen Wills Neuroscience Institute. His scientific focus has been on the use of molecular and cellular engineering technology to treat biomedical problems.
Most of the cells in the human body are specialists assigned to a specific organ or type of tissue, such as the neuronal cells that wire the brain and central nervous system. Stem cells are different. When they divide, they can produce either more stem cells, or they can serve as progenitors that differentiate, as they mature, into a specialized cell. Hence the name, because specialist cells can “stem” from them.
The potential to differentiate into specialist cells whose populations in the body have become critically depleted as the result of illness or injury is what makes stem cells so potentially valuable to medical research. If a batch of stem cells could be directed down specific pathways, they could be grown, harvested and then transplanted into a problem area. The plan is for these new cells to overcome damaged or diseased cells and lead to healing and recovery.
“The life of a stem cell can be viewed as a hierarchical branching process, where the cell is faced with a series of fate switches,” Schaffer says. “Our goal is to identify these cell fate switches, and then provide stem cells with the proper signals to guide them down a particular developmental trajectory.”
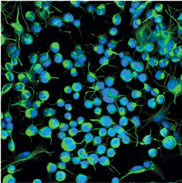 In the presence of the Sonic hedgehog protein, adult neural stem cells in culture began to proliferate. Those stained in green are on their way to becoming neurons.
In the presence of the Sonic hedgehog protein, adult neural stem cells in culture began to proliferate. Those stained in green are on their way to becoming neurons.
To date, most of the research and just about all of the publicity about stem cell research has focused on stem cells found in four-day-old human embryos, or blastocysts. These embryonic stem cells are prized for medical research because they have the potential to become just about any type of specialized cell in the body.
However, in some types of tissue there are pockets of adult stem cells that have the potential to differentiate into the cells that make up their host tissue. For example, adult neural stem cells residing in the hippocampus and in a region in the forebrain called the subventricular zone can generate new neurons. Working with these adult stem cells offers some advantages over embryonic stem cells, as Schaffer explains.
“First, adult stem cells don’t present the ethical dilemma that embryonic stem cells do because harvesting them does not destroy the donor. Also, they have already made some choices towards differentiating along a specific developmental pathway, so that makes the job of directing their future choices down that same pathway a little easier.”
The first hurdle in working with adult stem cells is getting these cells to proliferate. Normally, adult neural stem cells are present only in minute quantities, and most of those choose to remain quiescent.
“Adult neural stem cells don’t seem to exist for the purpose of replacing damaged neurons, but we believe by identifying and learning to manipulate the signals that control their proliferation and differentiation we can use them for that purpose,” Schaffer says.
Towards this end, he and his colleagues have taken both theoretical and computational approaches to unmasking the regulatory elements in adult neural stem cells. In their modeling work, they focused on a few prime suspects, such as the Sonic hedgehog protein. This soluble signaling protein was known to play a critical role in controlling the proliferation of numerous mammalian cell populations during early development.
To determine what role, if any, the Sonic hedgehog protein played in adult mammals, Schaffer and his colleagues harvested neural stem cells from rats, then cultured them with Sonic hedgehog protein. Their success in boosting the proliferation of adult neural stem cells in vitro led to in vivo experiments. Again working with adult rats, they delivered the Sonic hedgehog gene to the hippocampus of each experimental animal via a special virus. This resulted in the observed tripling of neural stem cells. In addition, injecting cyclopamine, a known pharmacological inhibitor of the Sonic hedgehog gene, significantly reduced neural stem cell proliferation in test rats.
“The next step is to identify signaling proteins that regulate differentiation in neural stem cells,” says Schaffer. “We’d also like to know more about the mechanism by which the Sonic hedgehog protein is regulating stem cell proliferation. Cells are exposed to a lot of different regulatory signals which they must correctly interpret to carry out functional decisions. How does this information processing system work?”
Schaffer cautions that it may be another 10 to 20 years before adult neural stem cells play any significant role in therapy.
“With these neural stem cells, I think we’re now at the stage that research with monoclonal antibodies was in the 1980s,” he says. “However, if we continue to make progress, the potential implications for medical research are enormous.”
Collaborating with Schaffer on this research were Karen Lai and Matthew Robertson, both with UC Berkeley’s Chemical Engineering Department and the Helen Wills Neuroscience Institute.
Conference on Women in Science Coming to Lab

Laura Chen, host and organizer of Women's Forum.
The role and influence of women in science and other technical fields will come under the microscope during the first Laboratory Women’s Forum to be held at Berkeley Lab. More than 100 people from Lawrence Berkeley, Lawrence Livermore, Los Alamos, and Sandia National Labor-atories will gather for two days on Oct. 26 and Oct. 27 in Building 66 to strategize about how the national labs can promote participation of women in technical fields and senior management.
“Inviting the widest possible points of view can have immense value in science and technology,” said Sally Benson, Berkeley Lab’s deputy director for Operations. “Gender diversity is one example of that breadth. Diversity in a field often increases its legitimacy and the value of its work in the public perception, and enhances the quality of decision making. I welcome the forum and encourage its efforts in achieving a more diverse workforce in technical and senior management at the national laboratories.”
Benson, who will welcome participants at the opening dinner, will be one of the many high-profile speakers who will address the event. Among them will be Lab Director Steve Chu, who will open the program the next morning; Congress-woman Ellen Tauscher; M.R.C. Greenwood, UC provost and senior vice president for academic affairs; Dick Nolan, head of the DOE Berkeley Site Office; and Deputy Lab Director Pier Oddone. Also scheduled to attend are UC President Robert Dynes and Bruce Darling, UC senior vice president of university affairs.
It is indeed noteworthy that the list above contains more men than women. But organizers are encouraged that the strong support from Lab, DOE and UC management will bring about the kind of change the forum hopes to achieve.
“I’m so pleased that so many dynamic men lend their support to a women’s forum,” said Laura Chen, host and organizer of the forum. “This shows the huge support and commitment for women in science and engineering. It’s very encouraging.”
Conference highlights will also include a panel discussion on the impact of science and technology on women’s issues and a poster session with displays from all four laboratories.
Chen worked long hours since the beginning of the year (many of them on her own time) to organize this event — an ambitious goal to which she committed herself after first participating in the 2002 Women’s Forum in Albuquerque, N.M.
“That was a fantastic forum, and it motivated me to try to bring it to our lab,” she says. “We need to bring awareness to these laboratories about women’s issues and the need to promote the participation of women in science, engineering, and other professional fields. Other labs have strong bodies representing women’s issues. I hope key managers and scientists will get together and develop something similar here.”
A major focus of the conference will be what’s referred to as the “pipeline” issue. Although more women than ever enroll in science and engineering programs at the university level, the percentage of women who successfully pursue these careers afterwards drops significantly. The afternoon pipeline session will explore how national labs can do a better job of attracting and retaining women in science and engineering.
“Women are given incentives in college to follow that career path, but after they graduate and enter career market, there’s very little encouragement to continue in the field,” Chen says. “I hope this forum will strategize on how to encourage women to continue to pursue these fields and mentor them here at Berkeley Lab.”
The first day of the conference will feature afternoon tours of the Lab and campus, followed by a welcome dinner. The next day will review the progress made since the two previous forums held in 2002 and 1999. Capping the two-day event will be a networking session at the Berkeley Art Museum.
For many participants, this will also be their first opportunity to visit Berkeley Lab, something that makes Laura Chen especially proud.
“From a personal standpoint, I also want to show off our lab,” she said. “I’m so proud of it. I’m very encouraged and confident that the forum will have many benefits to our laboratory and beyond in the long run.”
For more information about the forum, see http://fac.lbl.gov/LWF2004/ index.htm
Director Steve Chu�s First Talk to Employees Emphasizes Sense of Community at the Lab
Last week Director Steve Chu made his first official appearance before the Laboratory staff since arriving on Aug. 1. The 200-plus recipients of his introductory remarks got their first look at and first impressions of the person who will lead this Lab into the 21st century.
What they got was an uplifting, encouraging message, delivered casually and straight from the heart. With dashes of humor sprinkled in, Chu served a flavorful stew of promise, pride and optimism.

The crowd at the cafeteria enjoys a light-hearted moment during the
reception.
“I am delighted to be here,” he told them from his speaker’s platform in the cafeteria dining room. “This is really the greatest DOE lab there is, and we have good evidence for it. Like, for example, nine Nobel prizes — you should take great pride in this.”
He could have added his own Nobel as a tenth, but instead, he focused on those earned by Berkeley Lab scientists, promising “at least one more in the pipeline. It’s only a matter of time.”
Affable, quick with a smile, and precise with his words, Chu urged everyone in attendance to “feel you’re working toward a larger goal as a collective unit. There have been ‘golden moments’ in science — for example, Ernest Lawrence and the period of the Nobelists, that was their golden moment. Well, I want to make [today’s Lab] a golden moment in science, too. We have all the pieces here. Science is lots of little things cumulatively adding up to big things.”
In particular, he talked about the Lab’s broad experience in fundamental science, its success in responding to national research needs, and its reputation for operating some of the world’s most powerful user facilities. As an example of future pursuits, he cited the prospect of a post-fossil-fuels world and science’s contribution to it. “One of our greatest challenges is to get on a sustainable diet of energy that is CO2-neutral. Population will grow, and developing nations will develop. It will all take energy, and we don’t have a solution.”
Chu said it may all boil down to three options: fission, fusion or a solar derivative. He even speculated that plants, which convert solar to chemical energy, may hold first clues to a process that might be manufactured in a laboratory. “We have three to five divisions that could make an impact,” he told the audience. “It is a collective pursuit of a problem that no other labs can equal.

Director Chu chats with Armando Bautista and Mary Clary as he informally
meets employees following his talk.
“My role,” he continued, “is to try to create an environment where the best ideas will bubble up from the bottom, to build an (adequate) infrastructure, and to make sure Washington knows how good we are.”
He listed among his top priorities the strategic construction of new facilities that can replace buildings on the Hill that he said “were old when I was a graduate student here in the ‘70s. And they aren’t getting any younger.” He considers such capital upgrades crucial to sustaining the excellence of both the Laboratory and the UC Berkeley campus, which the former Stanford professor said educationally should be “as good or better” than private schools like Harvard and, yes, Stanford.
Upon arrival at the staff reception, he joked about his “handlers” getting him prepared for the occasion, and he introduced his primary handler, wife Jean, who watched from the side with her 90-year-old mother, Mrs. Lewis, a recent visitor from her native England.
Upon departure, he left his new constituents with an upbeat benediction: “Feel good about yourselves. I hope in the next five or ten years, you’ll feel good about me. In any event, we will go forward and sink or swim together.”
Runaround ‘04

Setting a personal Runaround record, Dula Parkinson crossed the finish line last Friday in just 9 minutes and 15 seconds (the fifth best time ever) to win the men’s race — for the third consecutive year! That’s a distinction unequalled by any male runner in the event’s 27-year history. Not to be outdone, however, the women can take equal pride in their top finisher, Jamie Baskin, who won the Runaround for the third time as well, although her previous wins happened in 1999 and 2001. Bascom’s time was 12 minutes and 15 seconds.


Lab Deputy Director Pier Oddone congratulates the winners in the men’s and women’s categories, Dula Parkinson (left) and Jamie Bascom (right).
While not everyone vied for such accolades, an impressive 755 walkers and runners of all ages crossed the finish line, the fourth largest number of participants in the event’s long and successful history. Participants ranged from toddlers to Oliver Morse, the oldest finisher at 71.
Runaround organizer Stephen Derenzo would like to stress that the event could not have happened without the generous assistance from the many individuals and committees that dedicated their time for many months, as well as the Employees Activities Association, which provided the t-shirts. The winning t-shirt design was the artful creation of Shannon Magee, the web and graphics manager for the Lab’s Center for Science and Engine-ering Education.
“It’s remarkable how well the committees served over the past few years,” Derenzo said. “We’re like a well-oiled machine now.”

Runaround finishers receive the most coveted prize of the event: the Runaround T-shirt, designed by Shannon Magee.
At the exact same time that participants on the Hill took off in the Runaround — 12 p.m. Pacific and 3 p.m. Eastern time — members of the Berkeley Lab’s Washington, D.C. Site Office ran around the Mall in the nation’s capital, and walked away with the same colorful t-shirts.
The full results of the Run-around, including the winners in each age category, everyone’s time, and past records will be available soon on the Runaround website at http://cfi.lbl.gov/~ derenzo/runaround/.

Steve Derenzo (right) congratulates the youngest participant in the Runaround
Snapshot Yields Inside Look at Molecular Movement
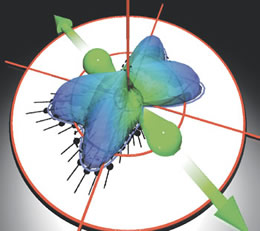
Black dots reveal the angular distribution of one electron, and the repelling nuclei are shaped like a green barbell in this snapshot of the breakup of a deuterium molecule. The second electron is emitted perpendicular to this plane.
The secret lives of molecules are now less secret. Using the Advanced Light Source, an international team of physicists has obtained the clearest snapshot yet of the simultaneous behavior of all the electrons and nuclei inside a molecule. Their work, in which they broke apart a deuterium molecule and measured the momenta of its particles, opens the door for a more basic understanding of molecules and the everyday processes they drive, from breathing to rust to photosynthesis.
“Nothing stands still. By learning how particles move, we can probe the fundamental properties of molecules and how they work,” says Thorsten Weber, a visiting scientist in Berkeley Lab’s Chemical Sciences Division who conducted the research with several other Berkeley Lab scientists, as well as physicists from Kansas State Univer-sity and institutions in Australia, Germany and Spain.
Like the opening shot of a pool game, the team fired a single photon at a two-atom deuterium molecule and broke it into its four charged constituents: two electrons and two nuclei, each containing one proton and one neutron. Next, the energy and direction of each particle was measured as it flew from the explosion and hit a position-sensitive detector. This information enabled the scientists to backtrack and determine how the particles were oriented inside the molecule precisely when the photon struck. The result is a multidimensional picture of the particles’ movement at the moment the molecule disintegrates, an image that inches physicists one step closer to directly observing the inner workings of a molecule.
“We obtained a fingerprint of the moving particles inside the molecule at the time of photo-ionization,” says Weber.
The combined momenta of a molecule’s electrons and nuclei dictate its geometry and how it binds with other molecules — in other words, what makes the molecule tick. But pinning down all of the particles’ momenta at the same time has proven extremely elusive. Theoreticians, relying on quantum mechanics, can only predict the probability that an electron will possess a given position or momentum.
To get an inside look at all the particles’ dynamics, experimental physicists are developing ways to fragment a molecule in a manner that preserves at least some of its internal motion, which is no easy task. Hit a molecule with an ion, for example, and the ion’s momentum transfers to the electrons and nuclei, which clouds physicists’ ability to determine their true momenta at the time of impact. But bombard a molecule with a photon, which has no mass and no charge, and the photon mainly deposits its energy and kick-starts the fragmentation process. With this in mind, the team used a photon from a polarized light beam generated at the Advanced Light Source to fragment a deuterium molecule.
Although physicists can’t yet peer inside a molecule without altering it, this “microscope for motion,” as Weber calls it, gives scientists the best vantage so far of the simultaneous momenta of all of a molecule’s particles. Its snapshots will enable scientists to someday peer inside more complex and biologically important molecules such as water and carbon dioxide, and observe how their particles govern life-sustaining reactions. The snapshots of the fragmentation of a deuterium molecule already give theoreticians something new to ponder.
“We experimentalists are ahead right now. Calculating such a few-particle break-up is a big challenge for state-of-the-art quantum mechanics,” says Weber.
Doing More with Less
Lab Scientist Develops New Way to Learn How Large Proteins Move

Paul Adams
In a development that will help scientists better determine how many large proteins work, Berkeley Lab’s Paul Adams and collaborators used the latest advances in computational analysis to study how a complex biological machine refolds proteins, a process critical to cell survival.
They took advantage of the fact that the protein, called a chaperonin, encapsulates unfolded proteins by moving large portions of its structure in unison, like a hand clenching a marble. This means that only a few frames of crystallographic images, each revealing the protein at a different stage of its motion, are needed to picture the entire process. And because many large proteins are believed to behave this way, their work also means that the mountain of crystallographic data collected over the past 10 years — much of which only portray large-scale changes in proteins — can help determine how many proteins twist and turn.
“We can get more out of crystallographic experiments than we thought we could,” says Adams, a staff scientist with Berkeley Lab’s Physical Biosciences Division and the deputy principal investigator of the Berkeley Structural Genomics Center. “Although we often can’t model the behavior of a protein atom by atom, it is possible to visualize how large portions of a protein’s atoms move together.”
As of early October, there were more than 27,000 structures in the Protein Data Bank, a worldwide repository of three-dimensional structures of large proteins and nucleic acids. Not all of these proteins lend themselves to this broad-brush analysis, because not all of them exhibit what’s called rigid body movement, in which discrete groups of their atoms move as a single unit. But for those that do, Adams’ technique could offer an efficient way to understand how they move.
His team, which included Charu Chaudhry and Arthur Horwich of Yale University and Axel Brunger of Stanford University, developed the method using a chaperonin found in Escherichia coli called GroEL. Like similar molecules found in the cells of all organisms, GroEL repairs proteins that unfold due to adverse conditions, such as heat stress. If too many proteins unfold, they cling together and cause cell damage, even death. To keep cells healthy, GroEL somehow surrounds a poorly folded protein, isolates it with a lid, then quickly stretches the protein before letting go, helping it move back to its correct shape.
To visualize how GroEL conducts this rescue work, the team used crystallographic data representing the protein in four different configurations: with its lid off, with its lid on, bound to the energy-providing enzyme adenosine triphosphate, and immediately after it attempts to refold a protein. They then ran this data through newly developed software that calculates how groups of atoms within the protein rotate and translate around fixed points. In this way, they were able to model GroEL’s movement using only about 20 parameters per rigid body, as opposed to the thousands of parameters needed to model a protein’s atom-by-atom movement.
“Our overall goal is to take crystallographic data and extract information about large motions in big systems,” says Adams.
New Way to Tackle Cancer Funded by the National Cancer Institute

Developing a mathematical model involves monitoring cultured cell-lines.
With a grant of more than $2.5 million per year for the next five years, the National Cancer Institute (NCI) has funded an Integrative Cancer Biology Program that Life Sciences Division Director Joe W. Gray characterizes as a “complicated, high-risk undertaking” aimed at predicting the success of new drugs that target cell-signaling pathways.
“This whole project builds from the fact that the private sector has developed some 800 pathway-specific therapeutic agents now undergoing clinical trials,” says Gray. “The cost to bring a new medicine to market is hundreds of millions of dollars, and the process takes decades. But only a small percentage of promising drugs ever get through the clinical trial stage.”
Co-principal investigator with Gray is Frank McCormick, director of the Comprehensive Cancer Center at UCSF, where Gray is also a professor. Among Berkeley Lab researchers, Mary Helen Barcellos-Hoff, Mina Bissell, Paul Spellman, Martha Stampfer, Damir Sudar, and Paul Yaswen of Life Sciences will be working with Jian Jin of Engineering and Victor Markowitz and Bahram Parvin of Computational Research, and with investigators at SRI International and the Nether-lands Cancer Institute.
Signaling pathways are inviting targets for drugs because their deregulation is implicated in many aspects of cancer. Among other functions, signaling pathways regulate how cells are organized, how they communicate with one another, how they proceed through the normal cell cycle, and when they undergo programmed cell death. Cancer-causing deregulation can occur at many levels, and tumors that appear clinically identical may have wide genomic variations.
“Many patients whose cancers appear to involve the same deregulated pathway don’t respond even to successful drugs,” Gray says. Ironically, in large trials patients who respond well to a particular drug may be so few that, statistically, the drug looks like a failure.
“We need to speed up the process of clinical trials and make it cheaper,” says Gray. “Once we determine exactly which factors are involved, we hope to predict who’s going to respond to a particular intervention, which will make it possible to do smaller trials.”
To this end, NCI has charged Gray and his colleagues with doing the experimental work to “tease out these factors and develop a predictive computer model.” To gain what Gray calls “a toe-hold on the problem,” with hundreds of signaling pathways already being targeted, the researchers are beginning with just one, a much-studied pathway known as Raf-MEK-ERK, which regulates the cell cycle and a host of other functions. Its misregulation is closely associated with breast cancer.
The backbone of the project is the large number of different breast-cancer cell lines that have been cultured at Berkeley Lab. Fifty-five such lines have been compared to cells from almost 150 human tumors and found to carry approximately the same genomic aberrations, making it possible to use the cultured cells to mimic the heterogeneity of real tumors in vitro. Growing, manipulating and monitoring the responses of so many lines is a challenge for the advanced robotics of Berkeley Lab’s Engineering Division.
“Managing data on such complicated molecular response profiles has never been attempted on this scale,” says Gray. Even a “simplified” Raf-MEK-ERK pathway diagram is as complicated as an integrated circuit, and testing responses to targeted therapeutics will be daunting. The collaborators will draw on the expertise in cancer-relevant pathways of Rene Bernards and his laboratory at the Netherlands Cancer Institute, while bioinformaticians at Berkeley Lab will be working with researchers at SRI International who have developed Pathway Logic models to tackle complex biological systems.
Gray emphasizes that developing a robust mathematical model of the Raf-MEK-ERK pathway is a formidable challenge; on the way to that goal, responses will be identified that can be used to predict who can benefit from specific therapeutic agents, and under what conditions. The tools developed for the task will be the first in an expanding kit that will hasten the debut of effective agents for fighting cancer.
Crystallization In Silico: Another Way To Image Big Biological Molecules
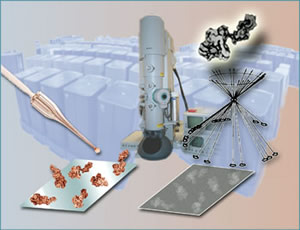
Macromolecules in solution are quick-frozen on a carbon stage. The electron microscope records 2-D images of the randomly oriented particles, and the computer recreates a 3-D model from a selection of these.
X-ray crystallography has long been the workhorse for imaging proteins and other biological macromolecules. But making crystals is tricky, and many proteins can’t be crystallized at all.
Enter single-particle electron cryo-microscopy. If making crystals is impractical, it’s possible to quick-freeze a sample and use an electron microscope to create 2-D images of thousands of randomly oriented “single particles” of the macromolecule.
“To see the structure in 3-D, you have to merge the data from all these individual images,” says Bob Glaeser of Berkeley Lab’s Life Sciences and Physical Biosciences Divisions and a professor of biochemistry and molecular biology at UC Berkeley. Once that’s done you have effectively constructed the crystal in the computer — what Glaeser calls “crystallization in silico.”
So far, single-particle methods have been restricted to resolutions of about 10 angstroms or lower, because “if you want atomic resolution, you’ll need a million particles or more,” Glaeser says. Today’s fastest computers would take 20,000 years to run the calculations.
Glaeser is co-PI with single-particle pioneer Pawel Penczek of the University of Texas in an NIH Program Project aimed at breaking the logjam. From its beginnings as a Berkeley Lab LDRD project, this high-resolution electron microscopy effort has involved the Computational Research Division’s Scientific Computing Group, headed by Esmond Ng.
Ng says that after he and group member Chao Yang learned single-particle methods from the experts, they realized that “certain computing tools, not new but not used by structural biologists” could be applied to simplify the calculations.
“The experimentalists work with specifics, from the bottom up,” says Yang. “We saw a top-down approach to the general problem.”
The result is a radical streamlining, reported by Yang, Ng, and Penczek in the forthcoming November 6 issue of the Journal of Structural Biology. Watch for details in the November edition of online Science Beat.
Lab CFO Jeffrey Fernandez Optimistic About Lab’s Financial Future
It was exactly one year ago that Jeffrey Fernandez, then deputy chief financial officer (CFO) for Lawrence Livermore National Lab, was summoned to Berkeley Lab to help straighten out a tottering financial structure. A three-to-four month stay, his consulting contract said. He had no intention of staying.
Today, Fernandez is Berkeley Lab’s CFO, his appointment coming in the wake of an impressive turnaround that persuaded management here that he was the one to lead the Lab full-time out of its fiscal system doldrums. His impressive professional resume supports the decision: 20 years at LLNL, with jobs that included deputy CFO, budget officer, and earth sciences associated department head; and earlier service with both the Executive Office of the President’s Office of Management and Budget (OMB) and the Depart-ment of Energy’s Office of the Controller.
As Berkeley Lab’s top financial executive, Fernandez has responsibility for about 180 employees and several departments, including the Controller and Budget Office, Procurement, Sponsored Projects, and Property Management, which will be transferred to the office this fall. In his fifth-floor office in Building 937 overlooking University Avenue, he talked to The View about his first year in Berkeley.
Jeffrey Fernandez
VIEW: You were brought to the Lab to engage in a major reconstruction of financial services here. What were the problems that needed to be fixed and the challenges you faced in fixing them?
FERNANDEZ: The problems were primarily in two areas. First, the people here were under a lot of stress. There had been a lot of employee exodus in prior years, leaving thin staffing levels. When financial problems arose, there weren’t enough people to work them. The second issue was external perceptions and lack of confidence in the ability of our lab to manage its own finances — things like just keeping the books up to date. People were overworked, and they had neither the bandwidth nor the time to keep up. And with all that, I came during the period of year-end close, right after several negative events, and people were working very hard and were discouraged.VIEW: What is the status of the office reorganization now, and what still needs to be done?
FERNANDEZ: It’s nearly complete. We’re trying to build one streamlined financial process, rebuilding both the financial system and the tools while integrating the field resource analysts and the central core. They (about 45 analysts in divisions and departments) used to be part of ASD (Administrative Services Division). Now all financial employees are within the Office of the CFO. This will allow us to train all financial professional employees to the same standard. This also helps with communication, and problems eventually become common rather than “theirs” and “ours.”
We identified a number of additional positions we needed, and we needed to expand our focus in areas like financial policies and training, rebuilding the Budget Office, and augmenting the staff in different accounting offices. We should be complete by the end of the calendar year.VIEW: What concepts and philosophies did you bring from Livermore to apply here, and what things did you find here that are uniquely Berkeley?
FERNANDEZ: Beyond the similarities inherent in scientific research facilities, I have really learned a lot since I’ve come here about profound differences between the two. They are mostly around the nature of the programs, the relationship to the (Berkeley) campus, and the integration with the university. The value structures, the dynamics, the cost drivers are different. What I brought with me were the importance of communications in all directions, the notion that the quality of the people drives how successful we can be, and the need to provide value-added service to the programs. The more involved we can be, the better service we can provide. If we can be included in discussing the programmatic objectives, we can help with strategies such as obtaining different types of funding for the different needs.VIEW: What lies ahead for Berkeley Lab’s budget? Good or challenging times?
FERNANDEZ: Probably both. This lab has identifiable national leadership in certain programs that will help it to survive through the projected flat budgets of the DOE — programs such as computing, life sciences, nanotechnology. In work for others, especially the NIH (National Institutes of Health), we have opportunities for significant growth.VIEW: Are there areas you will prioritize for improvement in the coming year?
FERNANDEZ: Number one, to streamline the financial process and the tools to make it easier for the customer to make decisions and get the mission done. And secondly, integrate all the financial professionals. That includes town hall meetings, a monthly meeting with the senior financial managers within each division, and opportunities we might have to socialize. For example, we have a Halloween party scheduled for the cafeteria.VIEW: What is it you liked about Berkeley Lab that persuaded you to come here?
FERNANDEZ: The people here were very welcoming, especially within the office of the CFO, and also the deputy directors and the division directors. I was favorably impressed with the Lab employees’ commitment to the Lab’s mission and to the science. Also, there are a lot of new things for me to learn here. And I’m absolutely optimistic about the Lab’s future. We’re at the epicenter where disciplines like biology, physics, chemistry and computing come together, a nexus of expertise that no other facility has at quite this level. It’s an exciting and challenging place to work, and I feel very energized.VIEW: When you’re not dealing with high finance, what do you like to do to get away from it all?
FERNANDEZ: Well, I have a 15-month-old son, and I like to wrestle on the floor with him while I can still pin him. I also cook for family and friends (he has attended culinary school) and drink fine wine. Also, my wife Nancie and I have strong ancestral ties to New England (Fernandez grew up there and graduated from the University of Vermont). We’re starting to build a timber-frame summer and retirement home on Echo Lake in the Northeast Kingdom of Vermont, with wood milled from the trees on the land. The nearest traffic signal is 30 miles away, there are loons on the lake, and the water is pure enough to drink. In late October, we’ll clear the lot and drill the well, and hopefully next June we’ll have a “raising day” (the day when the timber frame structures assembled on the ground are raised up and joined together). The families of the construction will all come and have a big party at the end of the day.
Berkeley Lab View
Published every two weeks by the Communications Department for the employees and retirees of Berkeley Lab.
Reid Edwards, Public Affairs Department head
Ron Kolb, Communications Department head
EDITOR
Monica Friedlander, 495-2248,
msfriedlander@lbl.gov
Associate EDITOR
Lyn Hunter, 486-4698 lhunter@lbl.gov
STAFF WRITERS
Dan Krotz, 486-4019
Paul Preuss, 486-6249
Lynn Yarris, 486-5375
CONTRIBUTING WRITERS
Jon Bashor, 486-5849
Allan Chen, 486-4210 David Gilbert, 925-296-5643
FLEA MARKET
486-5771, fleamarket@lbl.gov
Design & Illustration
Caitlin Youngquist, 486-4020
TEID Creative Services
Communications Department
MS 65, One Cyclotron Road, Berkeley CA 94720
(510) 486-5771
Fax: (510) 486-6641
Berkeley Lab is managed by the University of California for the U.S. Department of Energy.
Online Version
The full text and photographs of each edition of The View, as well as the Currents archive going back to 1994, are published online on the Berkeley Lab website under “Publications” in the A-Z Index. The site allows users to do searches of past articles.
Flea Market
- AUTOS & SUPPLIES
‘00 SUZUKI Grand Vitara JLX, hard top, 4 dr/wd, 2.5L/V6, at, cruise, 61K mi, exc cond, $7,600, Akira, X7296, 233-3665
HOUSING
ALBANY 2/2 condo on Park Hill, fully furn, 1,234 sq ft, sunny, quiet & eleg, laundry rm, views, pool, DSL, exer & rec rooms, 2-car garage, 848-1830, gfchew@ mindspring.com
BERKELEY, newly remodeled studio+ apt, 330 sq ft, downstairs in home, avail now, 10 min walk to LBNL, pub trans, priv ent, bay view, garden, kitchen, bth, all utils, cable TV, internet, $950/mo+occ’l pet/garden sitting. John, JLMcCarthy@ LBL.gov, 841-7875
BERKELEY, 1 bl to UCB, lge, nicely furn 1 bdrm apt, $1,575/mo incl computer, DSL, all util, Jin, 845-5959, jin.young@ juno.com
EL CERRITO, studio w/ spacious kitchen & bthrm, carpeted rm, lge walk-in closet, garage, 1 bl to the El Cerrito Pl BART, 6515 Central Ave, $875/mo Phyllis, 524-4849
EL CERRITO, 1 bdrm garden cottage w/ view, 1 bl to the bus, 7535 Stockton, $1,100/mo, Peter, 525-4406
NORTH BERKELEY by wk/mo, fully furn, 1/1 flat, quiet, spacious, dish TV, laundry rm, priv garden, gated carport, walk to shuttle/UCB/pub trans, Geoff, 848-1830, gfchew@mindspring.com
OAKLAND, sunny 1 bdrm in quiet triplex nr Rockridge shop ctr, hardwd flrs, lge din & liv rm, walkin closets, attached garage, close to trans to UCB/BART & downtown Oakland, $1,100/mo, Janice, 428-1893
PINOLE, 1 bdrm/1 bth duplex, 950 sq ft, lots of storage, lge priv courtyard, w/d, refrig, gas stove/oven, tiled throughout, berber carpeted in lge bdrm w/ tiled office nook, all util pd, cable HBO & Showtime, 1 offstr parking space, $1,150/mo, 410-1288
POINT RICHMOND, 1 bdrm & 2 bdrm apts in 8-plex, Pt. Richmond Village, spacious, laundry, offstr parking, extra storage, water/hot water/garbage incl, avail now, $900/mo for 1 bdrm & $1,100 for 2 bdrms, Evan, X6784
TARA HILLS, 2 bdrms in 4 bdrm/2 bth house, share w/ a female, $650/mo, avail 1/1/05, Liz, X2724, cell, 685-0005, Lbear2500@aol.com
HOUSING WANTED
LOCAL HOUSING NEEDED, out-of-town grandparents looking for housing for extended visit with grandbaby, furn or unfurn, 1 or 2 bdrms, mid-Dec to mid-Feb, pref within 5-10 mi of campus, will consider longer dist, bfisher@lbl.gov
MISC ITEMS FOR SALE
BICYCLE RACK, fits most sedans, mounts on trunk, easy to install, $15, Howard, X5031, 540-6718
BIKES, a Mongoose suitable for child ~ 6-8 & a nice mntn bike w/ fr shock, helmet avail, Fred, X4892
BOOKSHELF/DESK COMB, wooden bookshelf sits on top of desk, can be used sep, desk has 7 drawers, 3 on each side, 1 ctr drawer, like new cond, $100, Mary, 638-1451
CD/DVD PLAYER, Sony DVP NS300, silver, exc cond, orig box, manual, remote, cables, $45, Paolo, X6717
MIRROR, oval, beveled, gold leaf frame, 21x31, $25; beige lacquer dresser, 62”x29”x17”, $125; bronze ceiling fixture, faux alabaster glass, $25, Sue, 595-7123
TABLE FOR 4, extra leaf, leather chairs, good cond, b/o, Sue, X6432, 658-2426
TABLE, solid wood, round, seats 4+ & chair, $40 for both; twin size wood futton frame, $25; twin size mattress, almost new, $80; wood PC table, $5; blk wood night table, $10: floor lamp, $5, 14” col TV, no remote $10; 2-dr bthrm cabinet, fits under sink, $10, Cynthia, X4803, 486-9789
TRAINS, HO scale model engines, 4 ea, Atlas Gp38 No 7061 Union Pacific; 1 Spectrum GP-30 diesel; Great Northern 3000, 1 Athearn 3624 FP45 Pwr Amtrak; 2 Athearn 4502 SD40T-2 Pwr Cotton Belt; 12 cars; kits 1 Atlas passenger station kit #706; 1 model pwr water tank & shed #428; 1 model pwr “IRS on Fire” #470; $50 for everything, Mike, X5143, 525-4483
WANTED
BABYSITTER, new position for reliable babysitter, took exc care of twin girls for 2.5 yrs, Ulli X5347, 527-6643
VACATION
LAKE TAHOE house, 3 bdrm, 2.5 bth, fenced yard, quiet/sunny, great views, $195/night, 2 night min, Bob, (925) 376-2211
LAS VEGAS, time share, Polo Towers, 5-star in Las Vagas, 2 bdrms, 1 wk every other year or 1 bdrm, 1 wk every year, floating time, highest exch rating w/ resorts around the world, www.resortime.com, $17,000/neg, Mustafa, 451-1350, X5081.
SO LAKE TAHOE, spacious chalet in Tyrol area, close to Heavenly, full furn, sleeps 8, sunny deck, pool & spa in club house, close to casinos, attractions, $150/day+$75 1-time clean fee, Angela, X7712 or Pat or Maria, 724-9450
Flea Market Policy
Ads are accepted only from Berkeley Lab employees, retirees, and onsite DOE personnel. Only items of your own personal property may be offered for sale.
Submissions must include name, affiliation, extension, and home phone. Ads must be submitted in writing (e-mail: fleamarket@lbl.gov, fax: X6641), or mailed/delivered to Bldg. 65.
Ads run one issue only unless resubmitted, and are repeated only as space permits. Ads must be submitted in writing and be no more than 50 words in length. The submission deadline for the April 30 issue is Friday, April 23.

