
Two
Berkeley Lab Scientists Elected To National Academy
New Tools at DOE’s
Genomics Jamboree
Two Berkeley Lab Scientists Elected To National Academy
BY LYNN YARRIS
 |
 |
|
| Paul Alivisatos | Phil Colella |
Two Berkeley Lab scientists were among the 72 newly-elected members of the National Academy of Sciences (NAS) announced for this year. Paul Alivisatos, a chemist who serves as director of the Materials Sciences Division (MSD) and the Molecular Foundry, and Phil Colella, a mathematician who heads the Applied Numerical Algorithms group for the Computational Research Division (CRD), have joined the 1,949 active members of NAS. Established in 1863 under President Lincoln, the NAS is a private organization of scientists and engineers dedicated to “the furtherance of science and its use for the general welfare.” Membership in NAS is considered one of the nation’s most prestigious honors for a scientist or engineer.
Since joining the UC Berkeley faculty in 1988 as an assistant professor of chemistry, Alivisatos has steadily risen up the organizational ranks, while at the same time his scientific star has blazed. He joined the MSD staff in 1991, was promoted to associate professor in 1993, and became a full professor in 1995. An appointment as a professor of materials science and engineering was added to his C.V. in 1999, and he served as UC Berkeley’s Chancellor’s Professor from 1998 to 2001. In 2002 he was named to head Berkeley Lab’s Molecular Foundry, one of the proposed new U.S. Department of Energy centers for nanoscience research; and in January of 2003 he became the director of MSD.
During this period, as the leader of his own research group, Alivisatos scored a number of landmark successes in nanoscience, including the creation of the first semiconductor nanocrystals to be shaped like rods rather than spheres; the fabrication of a hybrid solar cell that advantageously combined nanotechnology with plastic electronics; and the co-invention of a cancer assay based on quantum dots. He has to date published more than 100 papers on the structural, thermodynamic, optical and electrical properties of nanocrystals.
Colella is a leading authority in the field of mathematics known as “adaptive mesh refinement.” In this technique, computer modeling algorithms are used to break large complex problems into smaller pieces in order to focus computing power on those areas of greatest scientific interest. His research group has developed advanced numerical algorithms and software applications in a wide variety of areas, including shock physics, turbulence, astrophysics, flow in porous media, and combustion.
Since coming to Berkeley Lab in 1996, Colella has won wide recognition for his work. Among the awards he has received are the 2003 SIAM/ ACM Prize in Computational Science and Engineering and the 1998 Sidney Fernbach Award from the IEEE Computer Society.
In addition to leading the Applied Numerical Algorithms Group, Colella also serves as project leader for the Applied Differential Equations Integrated Software Infrastructure Center (APDEC), sponsored by DOE’s Scientific Discovery through Advanced Computing (SciDAC) program, and for a project in NASA’s Computational Technologies for Earth and Space Sciences program called “Block-Structured Adaptive Mesh Refinement Methods for Multiphase Microgravity Flows and Star Formation.”
Both Alivisatos and Colella received their Ph.D.s from UC Berkeley. Alivisatos earned his degree in physical chemistry in 1988, and Colella earned his degree in applied mathematics in 1979. Colella was a professor in UC Berkeley’s Mechanical Engineering Department from 1989 to 1995.
New Tools at DOE’s Genomics Jamboree
BY DAVID GILBERT
 |
|
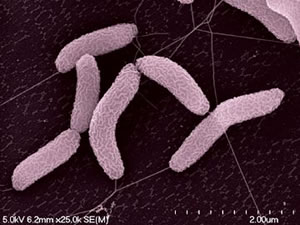
Although there were no s’mores munched around a campfire, those who participated in the DOE Genomics: GTL Jamboree at the Joint Genome Institute (JGI) last week did receive their just rewards. Armed with laptops and internet access, some 40 microbiologists, biochemists and computational scientists representing a dozen different labs, charted the unseen metabolic processes of one stinky “bug” that goes by the name of Desulfovibrio desulfuricans G20. Not a bug per se, G20 is a microbe with a robust appetite for such toxic metals as uranium and chromium, from a family known as sulfate-reducing bacteria, or SRB. Isolated from a corroded oil well in the late 1980s, the G20 strain was recently sequenced by the JGI.
Organized by Berkeley Lab’s Virtual Institute for Microbial Stress and Survival (VIMSS), the jamboree brought together scientists who clambered to compare and contrast SRB genomes, pitting G20 against other strains, including its cousin, D. vulgaris.
“This jamboree has crystallized a community,” says Adam Arkin, a Berkeley Lab scientist and director of VIMSS. “We have come together to see that sulfate reducers get their proper place in the biogeochemistry.”
 |
|
| Paramvir Dehal (left), Astrid Terry and Pilar Francio of JGI and Katherine Huang, a VIMSS software developer and scientist in Berkeley Lab's Physical Biosciences Division, examine genomic mysteries being unraveled during the Jamboree. | |
Sulfate-reducing bacteria are conspicuous because of the product of their respiration, hydrogen sulfide (H2S), smells of rotten eggs. Producing this extremely reactive gas — toxic to plants and animals — these bacteria thrive in anaerobic conditions of deep marine sediments, where oil can be found. Another idiosyncrasy — one that researchers are seeking to tap — is their generosity in donating electrons, a process known as reduction. In association with organics in the soil, SRB convert sulfate to sulfide — hence the big stink.
DOE’s interest in these microbes hinges on their ability to reduce uranium from its highly oxidized state. This reaction detoxifies the metal so that it is not as harmful to humans while making it insoluble in water, effectively immobilizing it in the soil.
There are significant economic and environmental pressures compelling the study of SRB, says Gerrit Voordouw, a professor of microbiology at the University of Calgary who chose Desulfovibrio as his bug of study back in 1983. “The oil industry wants to contain these bacteria because they cause souring, or production of H2S,” says Voordouw. The problem is exacerbated when drillers inject seawater, rich in sulfur, into the wells to force out the oil. Useful for liberating reserves, the practice, however, degrades oil quality and stimulates the bugs to corrode pipes, all revealed in the microbial physiology of down-hole conditions.
“As oil and gas become more depleted we have to think about alternatives, to harness photosynthesis for converting light into hydrogen. If you want to use biological pathways, you need to understand workhorse organisms like Desulfovibrio,” says Voordouw. “We can take it from there.”
Judy Wall is the keeper of the G20 flame. The University of Missouri, Columbia biochemistry professor is interested in metal metabolism and in enabling applications of SRB for bioremediation.
 |
|
| Desulfovibrio desulfuricans G20 |
“The more we know about how it generates energy, transports things, repairs DNA, responds to environmental assaults, the better we can make some reasonable predictions,” Wall says. These include horizontal gene transfer — a mechanism believed to be essential for genetic plasticity (the ability of bacteria to adapt to new environments).
Plumbing the depths of annotation data is where the toolbox comes in. Arkin’s group took inspiration from VISTA (Visualization Tools for Alignments), a winner of the Federal Laboratory Consortium Award for Excellence in Technology Transfer developed by a team led by Berkeley Lab’s Inna Dubchak. The new web-based tools on the VIMSS site (http://escalante.lbl.gov/) employ the “shopping cart” concept familiar to customers of such online retailers as Amazon.com.
Arkin calls upon the phone book analogy to explain. “Bob Smith in San Francisco and Bob Smith in New York have nothing in common except the name. But when we load all sequences we think are Bob Smith into the cart, then do analyses, we see that Bob Smith in Desulfovibrio vulgaris is likely the same Bob Smith that exists in E. coli.” He says that a hierarchy of function is established from which knowledge-based predictions can be made, including phylogenetic trees that show the evolutionary relationships between species.
 |
|
| Judy Wall of Missouri-Columbia (left) with Katherine Huang of Berkeley Lab and Xiangkai Li of the University of Oklahoma were among the 40 GTL Jamboree participants. In the background are Ralf Rabus of Max Planck Institute on the left and Chris Rao of Arkin Lab on the right. | |
Eric Alm of VIMSS highlighted a number of firsts among the jamboree’s findings, including the identification of a particular genomic signature of sulfate-reducing bacteria and several genes of unknown function that will be the target of future study.
“We identified DNA regulatory motifs and riboswitches — RNA sequences responsible for the synthesis of vitamins and other basic building blocks necessary for growth,” he said. “In addition, we identified a new regulon involved in the regulation of sulfate reduction.”
One of the major results was the first clearly resolved picture of the evolutionary history of the Proteo-bacteria, an ancient family of bacteria that includes the well-studied E. coli. The availability of whole genome sequences in the group known as Delta-proteobacteria including Geobacter metallireducens and Desulfovibrio desulfuricans G20 allowed us to build whole-genome phylogenetic trees revealing the details of the most ancient branches within the family tree for these organisms.” This work resulted from a collaboration with JGI’s own Paramvir Dehal and Pilar Francino.
Alm believes the jamboree was the first event to incorporate functional genomic data into the annotation process. Moreover, Arkin says, “we brought together people working on these organisms who don’t know each other. Something you can’t simply do in a conference call. It’s what makes this sort of large-scale science possible.”
OPA Winners in the Spotlight: Joe Eto Sheds Light on 2003 Blackout
BY ALLAN CHEN
 |
|
| Joe Eto |
When the lights went out in the northeast U.S. and Canada last August, Berkeley Lab’s Joe Eto was one of the experts the Department of Energy called on to help find out what went wrong. The blackout of 2003 affected tens of millions of people and caused billions in economic losses. Immediately, the two governments needed to find out what happened.
Eto, a scientist in the Environ-mental Energy Technologies Division, was appointed to the U.S.– Canada Power System Outage Task Force where, as a member of its Electric Systems Working Group, he played a large role in the initial fact-finding investigation and in the technical analysis of what wrong. The task force issued its final report this month.
For his work on this project, Eto has won an Outstanding Performance Award, an honor extended three times a year to Berkeley Lab’s best and brightest.
“As a researcher, participation in the blackout investigation was a vivid reminder of the urgent need for improved real time grid reliability management tools in order to keep the lights on and, consequently, of the importance of Berkeley Lab’s work in CERTS to address these needs,” says Eto.
 |
|
Eto is and has been in the thick of electricity grid R&D for some years now, managing and conducting research that will help improve the grid’s reliability. As program manager of the Consortium for Electrical Grid Reliability Technology Solutions (CERTS), he manages research that develops new methods, tools and technologies to improve the reliability of the U.S. electric power system and the functioning of competitive electricity markets. CERTS consists of 11 major universities, four national laboratories, including Berkeley Lab, and several private companies. It receives research funding from DOE and the California Energy Commission.
In his more than 20-year career at the Lab, Eto has published more than 140 papers on electricity reliability, transforming the marketplace for energy efficiency, integrated resource planning for utilities, and financing incentives for and evaluation of energy-efficient technologies and programs.
Harvard Holmes Prepares for a Soaring Retirement
BY JON BASHOR
After 34 years at Berkeley Lab, Harvard Holmes isn’t just retiring — he’s getting ready to take off.
 |
|
| Harvard Holmes at Truckee Airport in 2003. |
An avid flier who used to accompany his father as he flew around the state on surveying jobs, Holmes finally got his own pilot’s license in 1997 and bought an airplane a year later. But after his last day at the Lab on April 7, he packed his car and headed to Oregon. There, he and his wife Sara are taking a four-week workshop to get them started on building their own plane, a four-seat, pressurized Lancair model capable of cruising at 200+ mph. The project will take up to three years to complete, at which time, “I’m definitely going to hire a test pilot for the first flight,” Holmes said.
Holmes joined Berkeley Lab’s Applications Group in 1970, working on a graphics-editing program for the CDC 6600 computer. The work helped him earn his Ph.D. from UC Berkeley. Later he worked on a project to develop a demographic information system, funded by the Department of Labor. A high point of that work was that the project brought the first VAX computers to the Lab in 1979.
Later Holmes served as deputy and later as acting department head in Computing Services, providing UNIX logins, mass storage, and centralized computing. That work led him to joining a small team that he didn’t think had a very good chance of succeeding. The goal was to develop a winning proposal for relocating a DOE computing center at Berkeley Lab. The center was known as NERSC, and the rest is history.
When NERSC arrived in 1996, Holmes helped recruit new staff and then joined the transplanted Mass Storage Group, where he remained since.
“I’ve always had interesting things to work on,” Holmes says. “It’s been a fantastic career.”
Moving quickly between two points is one of the reasons Holmes likes to fly. In 2000, he and his wife took a three-week trip that took them to places such as Toronto, Bar Harbor, Cape Cod, Jacksonville, and the Grand Canyon. Their favorite destination, however, is Sedona, just four hours by “Air Harvard.”
In his new pursuit of building a plane, Holmes enjoys the support of his two grown daughters. One got her pilot’s license at 18 and is now an air traffic controller. The other is a successful financial planner. “She said we could afford it, so now we’re spending their inheritance.”
Berkeley Lab View
Published every two weeks by the Communications Department for the employees and retirees of Berkeley Lab.
Reid Edwards, Public Affairs Department head
Ron Kolb, Communications Department head
EDITOR
Monica Friedlander, 495-2248,
msfriedlander@lbl.gov
Associate EDITOR
Lyn Hunter, 486-4698 lhunter@lbl.gov
STAFF WRITERS
Dan Krotz, 486-4019
Paul Preuss, 486-6249
Lynn Yarris, 486-5375
CONTRIBUTING WRITERS
Jon Bashor, 486-5849
Allan Chen, 486-4210 David Gilbert, 925-296-5643
FLEA MARKET
486-5771, fleamarket@lbl.gov
Design & Illustration
Caitlin Youngquist, 486-4020
TEID Creative Services
Communications Department
MS 65, One Cyclotron Road, Berkeley CA 94720
(510) 486-5771
Fax: (510) 486-6641
Berkeley Lab is managed by the University of California for the U.S. Department of Energy.
Online Version
The full text and photographs of each edition of The View, as well as the Currents
archive going back to 1994, are published online on the Berkeley Lab website
under “Publications” in the A-Z Index. The site allows users to
do searches of past articles.
Tracking Parkinson’s Disease with PET Scan
BY DAN KROTZ
 |
|
| Jim O'Neil, head of Berkeley Lab's Biomedical Isotope Facility, with a remotely controlled synthetic unit used to make FMT and other radiopharmaceuticals. The development of automation technology is an important tool in radiopharmaceutical preparation because it minimizes radiation exposure and provides reliability and consistency in procedures involving high-energy, short-lived isotopes. | |
Berkeley Lab researchers are developing a way to use Positron Emission
Tomography (PET) scans to track the effectiveness of a gene therapy
that promises to help patients with severe Parkinson’s disease
control their symptoms without dangerous side effects.
Their work, which focuses on refining the use of a PET-detectable
chemical called a tracer, could allow scientists to evaluate how well
the therapy jumpstarts the brain’s ability to produce dopamine,
a neurotransmitter that diminishes in Parkinson’s patients.
Last month, in an important milestone, the scientists administered
the tracer to healthy people, and used PET to watch it accumulate
in a dopamine-producing region of the brain that grows quiet as Parkinson’s
progresses.
“We need a way to see what’s going on in the brain, and the PET tracer lets us do that,” says Jamie Eberling of Berkeley Lab’s Center for Functional Imaging, who is conducting the research along with fellow Berkeley Lab scientist Henry VanBrocklin. Other collaborators include Krys Bankiewicz, a professor of neurological surgery at UC San Francisco, and Bill Jagust, head of the Life Sciences Division’s neuroimaging program, who began the research with Bankiewicz in the early 1990s.
The ability to pinpoint the brain’s dopamine-production centers
could give researchers a quick and accurate way to gauge the gene
therapy’s success once clinical trials begin. Ultimately, the
tracer could speed the development of a better way to treat the advanced
cases of a disease that afflicts two million people in the United
States and Europe.
They chose PET because of its unique ability to detect the metabolic
and biochemical signatures of disease, which occur long before a disease
causes anatomical changes and clinical symptoms. Other imaging techniques
such as Computed Tomo-graphy (CT) and Magnetic Resonance Imaging (MRI)
can only detect structural changes.
“PET can be used to more quickly determine how the therapy is working,” says VanBrocklin. “There’s no better way to track the therapy in patients.”
The tracer, fluorine methyl tyrosine, or FMT, was developed at the
University of California at Los Angeles in the late 1980s. It’s
engineered to cross the highly selective blood brain barrier and enter
only those neurons that contain an enzyme that converts dopa to dopamine.
In Parkinson’s, for reasons that are not fully understood, neurons
that contain this enzyme decrease in number and the brain’s
dopamine levels dip precipitously — a phenomenon that leads
to the tremors and muscular rigidity that mark the disease.
Because dopamine-producing neurons reside in very specific regions
of the brain, a PET scan of a healthy person given the FMT tracer
should yield an image with bright spots, indicating areas where the
tracer accumulates. Conversely, a PET scan of a Parkinson’s
patient should reveal darker areas, indicating an absence of FMT and
an absence of dopamine-producing neurons.
For more than 10 years, Berkeley Lab researchers have worked to optimize the tracer’s ability to zero in on these dopamine-producing hotspots. They’ve even invented an automated robot that quickly synthesizes large quantities of the chemical. Now, after extensive lab work, they’re assessing how the tracer accumulates in the brains of healthy people between 40 and 60 years old, the age at which Parkinson’s symptoms first appear.
 |
|
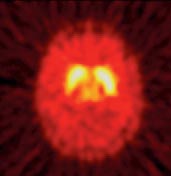
| PET image of the brain of a healthy person given the FMT tracer. As expected, FMT accumulation is highest in the striatum as shown by the more intense PET signal, with relatively little FMT uptake in other brain regions. |
On a separate track, researchers from the Bay Area-based company Avigen have created a gene therapy that’s designed to replace the dopamine-producing enzyme that disappears in Parkinson’s patients. It works by delivering the gene that produces the enzyme directly to the striatum, the area of the brain that needs dopamine to control movement. Although this therapy is not intended to alleviate symptoms, it may allow patients with severe Parkinson’s to take much smaller doses of the drug levodopa, the primary treatment for the disease. The drug, which is converted by the neurons’ dopamine-producing enzymes into dopamine, successfully controls tremors in the early stages of the disease. But as the disease progresses, and the level of dopamine-producing enzymes drops, larger doses of levodopa are required to suppress symptoms.
Unfortunately, large doses of levodopa have severe side affects such as uncontrolled flailing, psychosis and hallucinations. Avigen researchers hope their gene therapy will replenish the enzymes lost in patients with advanced Parkinson’s, enabling them to take safer doses of levodopa.
When Avigen begins evaluating the gene therapy in clinical trials, slated to take place at UCSF, they’ll use the PET tracer developed at UCLA and refined at Berkeley Lab. Ideally, in a series of PET scans taken over several months, a Parkinson’s patient’s striatum will grow brighter and brighter as the gene therapy provides it with a steady supply of dopamine-producing enzymes.
“The tracer is an excellent measure of gene expression,” says Eberling. “It will allow researchers to follow patients throughout the course of therapy and make sure gene expression is sustained.”
Ultralow-field NMR and MRI Now Available for Licensing
BY PAUL PREUSS
 |
|
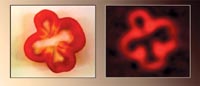
| A slice of red pepper (photo, top) was imaged with the new SQUID MRI system using a magnetic field of only 0.13 millitesla. | |
The Technology Transfer Department has announced that ultrasensitive systems for magnetic resonance imaging (MRI) are available for licensing. Pioneered by the Materials Sciences Division’s John Clarke and Alexander Pines and their colleagues, the systems use magnetic fields 10,000 times weaker than conventional MRI and its parent technology, nuclear magnetic resonance (NMR).
Conventional MRI typically requires a magnetic field of 1.5 teslas — roughly 30 times stronger than the Earth’s magnetic field. The systems developed by Clarke and Pines operate at a mere 1 to 100 microteslas (millionths of a tesla). Generating low-strength fields is far less expensive, and they need not be highly homogeneous to achieve good resolution.
The Clarke-Pines breakthrough first prepolarizes sample nuclei (typically protons) to increase signal strength, then detects the signals in an ultralow field with a superconducting quantum interference device, or SQUID.
Traditional NMR detects the frequency at which spinning nuclei precess; the stronger the magnetic field the higher the frequency, and the better the signal-to-noise ratio. But SQUIDs detect the nuclei’s magnetic signal directly, regardless of frequency. In fact, the weaker the field the sharper the signal: the Berkeley Lab researchers obtained narrow NMR lines from protons in liquids even in inhomogeneous 100-microtesla fields.
Standard NMR and MRI polarize the sample nuclei and detect their signals in the same high field. In the new method the sample is prepolarized in a field of 100 to 300 millitesla (thousandths of a tesla), which is removed before the image is acquired in a much lower, microtesla field.
To construct images, MRI imposes secondary fields whose strength falls off smoothly. In this gradient the linewidth of the NMR signal determines resolution. Narrow linewidth signals detected by SQUIDs allow MRI with high resolution in gradient fields three orders of magnitude smaller than the gradients used in conventional high-field MRI.
Berkeley Forum Focuses on UC Labs
BY LYNN YARRIS
 |
|
| Pier Oddone |
Berkeley Lab Deputy Director for Scientific Programs Pier Oddone spoke eloquently, but a Berkeley public forum on “whether the University of California should continue to manage the DOE national labs” would have been more aptly billed as whether UC should continue to manage the “weapons labs.” Of the three national laboratories that UC currently manages for DOE — Lawrence Berkeley, Lawrence Livermore, and Los Alamos — LLNL and LANL were the focus of attention and contention.
“I would hope there is no wide concern about UC’s continued management of Berkeley Lab,” said Oddone in his opening remarks to about 100 attendees at the Inter-national House auditorium where the forum was held on the evening of April 21. “We bring many UC professors and students to our Lab to work on problems of scale … and draw from some of the best campus minds. Berkeley Lab would not be what it is today without its close relationship with UC.”
The forum, hosted by KQED radio host Michael Krasny, was divided into two panel discussions. The first, in which Oddone participated, looked at the history and current status of the UC-DOE affiliation. Joining Oddone on that panel were Todd Haynes, a scientist at LANL, Michael May, director of LLNL from 1961 to 1975, UC Irvine emeritus history professor Karl Hufbauer, and Robert Powell, a political science professor at UC Berkeley. The second panel discussed the impact of the UC-DOE affiliation. Its members included Andrew Lichterman of the Western States Legal Foundation, Laura Nader, a UC Berkeley anthropology professor, William Kastenberg, a UC Berkeley nuclear engineering professor, and Marylia Kelley, executive director of TriValley CARES (Communities Against a Radioactive Environment).
Even the two most vocal opponents of UC’s continued affiliation with LLNL and LANL, Lichterman and Kelley, voiced support for the nonclassified “civilian scientific research” being conducted at Berkeley Lab. Haynes and May expressed their desire for continued UC management of their respective laboratories. Haynes said that “science is extremely valued” at LANL, and that while other potential management candidates, such as the University of Texas, might do a good job, “no other institute or organization compares to UC when it comes to doing science.”
Those advocating that UC discontinue its management of the weapons labs received the most audience applause. However, Kastenberg drew a lot of nods when he posed the question that if the U.S. chooses to remain a nuclear weapons nation, who better to be involved in the management and development of those weapons than UC?
“Is there anyone here who would rather have the responsibility in the hands of private industry?” he asked.
This forum was presented by the Berkeley Graduate Assembly and cosponsored by the Chancellor’s Committee for Dialogue of National and International Issues and the Institute of Nuclear and Particle Astrophysics and Cosmology.
Asf1: A Protein in the Thick of Things
BY PAUL PREUSS
 |
|
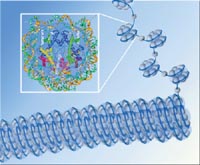
| Asf1 helps build nucleosomes, which are formed from eight histones looped with double-stranded DNA (inset). Strung together like beads on a string, the nucleosomes wrap themselves into chromatin, the stuff of chromosomes. | |
Picture a clipper ship with hundreds of sailors swarming over its rigging, each with a specific task but all working together to keep the ship safely under way. It’s not too different from the way a crew of proteins cooperates to keep cells afloat.
Among other tasks, these gangs of proteins build scaffolding on which DNA winds itself to make chromatin, the stuff of chromosomes; they open up double-stranded DNA so genes can be transcribed, or seal the strands to silence genes; they man checkpoints that regulate cell reproduction; they repair DNA when it breaks.
“One protein that’s found at the nexus of many of these
processes is a small, compact protein known as Asf1,” says Paul
Kaufman of the Life Sciences Division, who’s also an associate
adjunct professor in UC Berkeley’s Cell and Molecular Biology
Department.
Asf1 (antisilencing factor 1) is a “histone chaperone”;
its main job involves depositing histone proteins on DNA to form nucleosomes,
the linked protein-DNA units that make up chromatin. Asf1 also involves
itself in other critical functions like gene regulation and DNA repair.
The Asf1 gene was first identified in baker’s yeast, but, says Kaufman, “all eukaryotes have at least one version of the gene, and some, including humans, have two.” (Eukaryotes’ cell nuclei are contained in membranes.) To investigate Asf1 Kaufman joined forces with James Berger of the Physical Bio-sciences Division, associate professor of biochemistry and molecular biology at UC Berkeley, and collaborators from the Fox Chase Cancer Center in Philadelphia and Rutgers University.
“We were particularly interested in isolating the ‘core’ of this protein, the part that has been conserved during evolution,” Kaufman says. Asf1’s first 155 amino-acid residues are highly conserved in virtually all organisms, some separated by hundreds of millions of years of evolution; the rest of the sequence varies widely and may be missing altogether.
 |
|
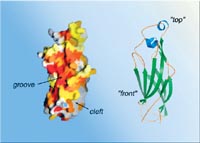
| A high-resolution model of the Asf1 protein’s core segment, using data from the Advanced Light Source’s beamline 8.3.1, reveals a number of likely binding sites where this versatile protein interacts with others. |
Kaufman and his colleagues used recombinant DNA technology to synthesize the core fragment: “We let evolution tell us where to cut their tails off.” The core fragments performed all the functions of the entire protein, binding histones and interacting with a cell-cycle-checkpoint enzyme in vitro, and in vivo resisting DNA-damaging agents in mutant yeast cells that lacked Asf1. The fragments interacted with other important chromatin-building proteins just like whole Asf1.
“The core of Asf1 is definitely the business end of the protein,” says Kaufman. To solve its structure Sally Daganzo, a member of his lab now at UCSF, grew crystals for x-ray crystallography with help from Berger’s lab. Using beamline 8.3.1 at the Advanced Light Source, the team constructed a 3-D model with a resolution of 1.5 angstroms, one and a half ten-billionths of a meter.
The researchers examined the model searching for “anything that looked like a docking site,” says Kaufman. Three regions stood out: a groove on the front of the molecule, a deep pocket, and an area of strong negative electric charge.
At one of these sites, Asf1 binds to the Hir protein in yeast and related HIRA proteins in humans, important in forming the parts of chromosomes inactive during gene expression. “Hir and HIRA continually deposit histones as they are building chromatin,” says Kaufman. “Asf1 may be the carrier that reloads their magazines.”
Kaufman emphasizes “We are in the process of looking carefully into how this one protein manages to find itself a central player in so many different processes. The full impact of what we know hasn’t come out yet.”
“Structure and function of the conserved core of histone deposition protein Asf1,” by Sally Daganzo, Jan Erzberger, Wendy Lam, Emmanuel Skordalakes, Rugang Zhang, Alexa Franco, Steven Brill, Peter Adams, James Berger, and Paul Kaufman, appeared in the Dec. 16, 2003 issue of Current Biology.
David Kaczynski Speaks at Lab About His Ethical Dilemma
BY DAN KROTZ
 |
|
| David Kaczynski | |
In 1995, David Kaczynski spent several sleepless nights wondering the unthinkable: was his older brother the Unabomber, responsible for 16 bombings that took place over 17 years, leaving three people dead and 23 injured?
After reading the 35,000-word anti-technology manifesto published in the Washington Post — which echoed remarks his brother had made in the past — and talking it over with his wife, David slowly grew unsure of his brother’s innocence.
“I was reading and reading, looking for the magic words saying Ted didn’t write this, and I couldn’t find them,” David Kaczynski said on April 14 in a presentation to a packed audience in the Building 50 auditorium. His talk, entitled “Doing the Right Thing — When It’s the Hardest Thing
to Do,” wasn’t a lecture, but rather a painfully honest account of his decision to turn his brother in, which ended one of the largest manhunts in FBI history.
“It’s a little sobering to be here. Just a few moments ago we drove by Cory Hall, where Ted placed two bombs,” David said. Although he was sure his decision was the right thing to do, and could possibly save lives, he admits it wasn’t easy.
“It was such a painful feeling to point on the map and tell the FBI where Ted lived,” he said. “But if we failed to act, and somebody died, my life would be meaningless.”
Ted Kaczynski was arrested several weeks later, and in 1998 received four life sentences without the possibility of parole. Although few people will be faced with ethical quandaries as overwhelming as turning in a relative, David concluded that such decisions should ultimately be based on a simple philosophy: “We need to create a culture where people care about each other. Our loyalty must be to humanity,” he said.
Earth Month at Berkeley Lab
 |
||
| Boise representative Gary Begg explains a new recycled paper shipping box to (from right) Beverly Harris, Robbie Jackson, and Kim Magbie, while Nancy Tallarico and Melanie Yip browse. |
The message of conservation was brought to the Lab last week in many forms. More than 20 groups educated employees at an Environmental Fair on Friday.
Did you know that an owl's eyes are fixed so they can't move from
side to side? To look at something off to the side, an owl has to
move its entire head, something it can do thanks to specialized neck
bones and muscles. With 14 bones in its neck, the owl can move its
head rapidly 270 degrees (three-fourths of a circle) to either side.
 |
 |
|
| Wildlife preservation was the theme of a lecture by Patti Blasquez (left) of the Lindsay Museum in Walnut Creek, who brought two of her feathered friends, a Swainson’s hawk and above right, a barred (due to its bar-like coloring) owl. | ||
Remembering Bill Hess: From Berkeley Lab to Apollo 11
BY PAMELA PATTERSON
 |
|
| Bill Hess (center) looks over the shoulder of Tom Seliga, director of the National Center for Atmospheric Research, as other NCAR staff members look on (1983). | |
Wilmot “Bill” Hess, a former Berkeley Lab physicist whose eclectic career took him from the halls of UC Berkeley to running the science program for the Apollo 11 moon mission, and finally to heading DOE’s High Energy and Nuclear Physics program, died in his Berkeley home on April 16 of leukemia. He was 78.
The son of a biologist — a discipline the younger Hess deemed to be “not structured enough” for his taste — he received his Ph.D. in physics from UC Berkeley in 1954. Like many newly minted physicists of that era, he immediately came up the Hill, running experiments on the 184-Inch Cyclotron.
“When I first walked into the building where the machine was, I literally sort of trembled and felt awestruck,” he would later recall. “I was the last generation of people who were able to work on individual experiments. The research team would think up the experiment, decide it was worth doing, develop the instruments to do it, carry out the experimental runs here in Berkeley, then analyze the data and publish the results. I liked that you could do the whole thing.”
Hess next migrated to the Lawrence Livermore Lab, working in space-related research, and then went on to Goddard Space Center in Maryland to become head of theoretical physics. It was a title he would wear with some discomfort for six years. “I was an experimentalist,” he said, “and after a few years of being head of the theoretical division, sort of a fraudulent title, I decided, I probably ought to go do something else.”
In 1967, lured by the chance to do experimental physics in the expanding frontiers of space, Hess took the job as director of science at the Manned Space Center in Houston for Apollo 11. His responsibility was designing the science for the Lunar Receiving Laboratory, the facility used to conduct scientific and engineering tasks. Recounting those early, heady days of the space race, Hess said the rules were simple: scientists could take 250 pounds of equipment and instruments down to the lunar surface, and bring back one hundred pounds — mostly consisting of lunar rocks and dust and the weight of the aluminum box they would be carted in.
Science, he admitted, wasn’t the foremost goal of the early Apollo flights. “President Kennedy sent us the ground rules: ‘Get a man there, get him back, and do a little science on the side if you can.’” Nevertheless, Hess considered getting the science done on Apollo one of his biggest accomplishments.
He recalled the moment Apollo 11 landed on the moon with typical boyish enthusiasm: “I was hearing the conversation between mission control and the astronaut [Neil Armstrong]. And the mission control guys said, ‘It’s going sideways. What’s it doing that for? Where’s it going? It’s only got forty seconds of fuel left!’ And they were very nervous. And then he [Armstrong] puts it down when he’s only got about twenty seconds of fuel left … and Oh, Oh boy!”
Always looking for a new challenge, Hess left the space program to become the director of the research laboratories of the National Oceanic and Atmospheric Administration in Boulder, Colo., where he worked for six years. Then, in his final venture, he returned at last to his roots as a physicist to oversee the energy labs, including Berkeley Lab, as associate director of High Energy and Nuclear Physics for DOE in Germantown. There he directed the research side of the ill-fated Superconducting Supercollider. Of that project, which had ultimately grown to $8 billion before being killed by Congress, Hess said, with characteristic candor, it was “just too much money.”
Hess retired from government service and returned to Berkeley in 1996. When an interviewer pointed out to him that his career took an interesting path, from physics to space science to meteorology and back to physics, Hess laughed, “Yes, I’ve had an interesting life. I wandered all over. Couldn’t hold a job.”
He is survived by his wife of 53 years, Winifred Lowdermilk Hess; a son, Carl Hess; and daughter, Alison Williams, as well as four grandchildren. The memorial service will be private.
Flea Market
- AUTOS & SUPPLIES
‘01 PRIUS, dark green, 19K mi, maintained, $15,000, Phila, 848-9156
‘00 SAAB 9 5 SE, 62K mi, very clean, $15,500, Vlad or Linda, 849-1579
‘97 WINNEBAGO Rialta TB, VW Eurovan V6, 201 hp, 69K mi, great cond, 3k generator, microwave, propane stove, roof, ac, tape player, telescope toilet & shower, twin beds, drives great, $23,000, Barbara, 652-7044
‘85 HONDA ACCORD LX, at, ac, 4 dr, pwr everything, 115K mi, rebuilt eng/trans/carburator, $1,500/bo, Bill, X7512, 813-1973
HOUSING
ALBANY, 1 bdrm condo, 555 Pierce St, unfurn, very clean, new carpet, exc cond, all util pd, garage, wheelchair access, bay view, storage, pool, tennis, gym, coin laundry, 24-hr sec, refrig, stove, dw, 12-mo lease, $1,100/mo + $1,500 sec dep, Mary, X4701, 816-9702
ALBANY, lge 2 bdrm/2 bth condo, fully furn, quiet & eleg, sleeps 4, laundry rm, views, swim pool, exer rm, rec rm, 2-car garage, close to food mkt/El Cerrito Plaza/ BART/trans, Geoff, 848-1830, gfchew@ mindspring.com
BERKELEY, Carleton & Grant, nr BART/ shops/Lab shuttle, close to LBNL, newly renov 2 bdrm apt, gr flr of 2-story Victorian house, sunny, front garden, w/d, custom tile flrs, no smok/pets, $1,400/mo, incl part utils, avail immed, Richard or Hope, 845-1723
BERKELEY, resid community of scientists/personnel/grad students, Hearst Commons, 1146-1160 Hearst, nr UC/pub trans/bike path, parking, studio cottages w/ decks, some furn can be provided, hdwd flrs, skylights, dw, ac, sec, lge studios w/ sleeping lofts & eat-in kitchens, $950, Ian, 220-2051
BERKELEY, avail 5/1, 2 bdrm/1 bth apt, partially furn, fp, DSL & cable ready, parking, laundry facil, nr Lab shuttle/UCB/shops, water & garbage incl, $1,700/mo, Christina, 525-6527
BERKELEY, clean, sunny rm in duplex, avail 5/1 for semester or longer, walk to UCB/Lab shuttle/ BART/buses/shopping, own phone, furn avail, $500/mo incl util, Lisa, 665-3146, stewartl@lmi.net
BERKELEY, furn rm w/ fast internet access, priv phone line, priv entry, quiet, safe, park, tennis court, swim pool, nr
BART/shops, offstr park, washer, gentleman, no smok/pets, no overnight guests, $600 + internet, dep $650, Ming, 524-3780
EMERYVILLE/OAKLAND, Berkeley border, furn bdrm in 2 bdrm/1 bth dup, private, lge double closet, garage, w/d, dw, furn liv rm, hardwd flrs, nr shopping/BART, share w/ female, sublet for 6 mo starting 7/04, dates neg, $550/mo + util, female pref, Prisca, 652-6434, pnodora@earthlink.net
MONTCLAIR/OAKLAND HILLS, house, 2 bdrm/1 bth house, priv, trees, decks, fp, laundry, easy drive to Lab, $1,950/mo, avail now, Fred, 486-4892
NORTH BERKELEY HILLS, furn home on Euclid nr Cragmont, avail 5/12 – 7/20, 2 bdrm/2 bth, den w/ hide-a-bed, view, secl backyrd w/ hot tub, garage, DSL, no smok/ pets, $500/week incl util, 6-week min, Steve, X5396, 813-1426, swiel@lbl.gov
NORTH BERKELEY B&B, darling cottage, 1 person only, $650/mo, breakfast daily, avail now, Helen, 527-3252
NORTH BERKELEY, rm w/ priv bth in quiet house, no smok/pets, furn w/ bed, desk, dresser, incl kitchen & laundry privil, nr pub trans, for single, responsible person, $500/mo incl util, Ms. Wurth, 527-5505 after 1:30 pm, nonet@xoma.com
NORTH BERKELEY, huge 1 bdrm house, fully furn, w/d, DSL wireless connection, no pets/ smok, $1,500/mo + dep + util or $600/wk, avail 6/7-7/20, shared rear & fr garden, 5 min walk to N. Berkeley BART, less than 2 mi from campus, Gilad, 219-4876, X5589
PINOLE, 3 bdrm/2 bth, exc cond, pet-friendly home, fenced yard, dog run w/ garage access, nr Pinole Valley Park, hot tub, w/d, fp, 2-car garage, view, great landscaping, easy commute, $1,800/mo, Erik, 910-4318, erpage@lbl.gov
RICHMOND HEIGHTS, housemate wanted to share beautiful 2 bdrm/2 bth, fully furn home in quiet neighbrhd, view, deck, fp, laundry, nr pub trans/UCB, pref female, non-smoker, $700/mo + $700 dep, avail now, short or long term neg, Joanne or Nan, 234-1064
ROCKRIDGE, by wk/mo, lge bdrm in furn, quiet & spacious 2/1 flat shared w/ owner in dup, priv deck, cable TV/VCR, laundry, garden & patio behind house, 1.5 blocks to Lab shuttle at Rockridge BART, public trans, Barbara, 652-7044
SAN FRANCISCO, rm for rent in 3 bdrm girl house, hardwd flrs, laundry, 2 full bths, huge kitchen & liv rm, sunny patio, nr BART & MUNI, $860/mo, 6/1, Daniela X5077, drusso@lbl.gov
HOUSING WANTED
HOUSESITTER or cat/dog sitter, for 1 wk any time this summer, will take care of plants & any pets, in Berkeley, Athena, 841-0849
LOST & FOUND
FOUND: TOTE BAG, black leather w/gym clothing, in Bldg 50B, 4th flr copy/break rm, Security, X4855
MISC ITEMS FOR SALE
COAT, women’s full length, bl leather, $65, Stephanie, X7058
COMPUTER MONITOR, 17” Optiquest V95 w/cable, exc cond, $30/bo, Pierre or Margot, 524-8672
CRATE FOR DOG OR CAT, approx 33x 22x26, $20/bo, Bob, 642-2156, 527-2937 eves
FILE CABINET, 2 drawer, top opens & holds file folders, very deep, on wheels, metal, will deliver to Lab, Diana, X4070
MICROWAVE, 1-yr old, white Panasonic, 1300 watts, $50; Panasonic VCR black, 4 head, perf cond, $30; Fujitsu life book laptop, Pentium II 233 Mhz/256 MB/3.2 GB HD, swappable CD-ROM dr, swappable floppy disk dr, USB port, 14.1” act matrix display, Win 98, PC ethernet card, bo; Linksys EtherFast cable/DSL router w/ 4 port switch, $25; Sony CD/radio/cass recorder, bo, Guoying, X5843
NIGHT STANDS (2), light oak, exc cond, $50 for pair/bo, Ken, X7739
PIANO, Wurlitzer console, 20-yrs old, exc cond, med walnut finish, good beginner’s piano, $800/bo, Ted, X4203, 548-9235
SOFA, 7 piece set; Whirlpool washer & dryer, oak wood din table w/ 4 chairs; queen & twin mattresses w/ boxes, 1-yr old, see @movingsale.8m.com, Sandhya, X5641
WANTED
HOUSESITTER, 1 wk, Sat, 5/15-22, no pay but very nice house in Oakland nr Hwy 13, care for 2 cats, Ken, X7739
VACATION & OUT OF TOWN
NORTH LAKE TAHOE house at Kings Beach, 3 bdrm/2 bth, sleeps 6, quiet cul de sac, 1 mi from pub beach, $135/night, 2 nights min + $75 clean fee or $675/wk, Vlad or Linda, 849-1579
WASHINGTON DC, 1 bdrm, fully furn condo in old town Alexandria, 1 min walk to metro, w&d, ac/dw, pool & weight rm, avail 5/1-8/1, $1,200/mo, util except electricity, Doug, 486-6095, cell 332-4097, dhale@lbl.gov
Flea Market Policy
Ads are accepted only from Berkeley Lab employees, retirees, and onsite DOE personnel. Only items of your own personal property may be offered for sale.
Submissions must include name, affiliation, extension, and home phone. Ads must be submitted in writing
(e-mail: fleamarket@ lbl.gov, fax: X6641,) or mailed/delivered to Bldg. 65.
Ads run one issue only unless resubmitted, and are repeated only as space permits. Ads must be submitted in writing and be no more than 50 words in length. The submission deadline for the April 30 issue is Friday, April 23.
What Do Scientists Do?
Kids Found Out During ‘Daughters and Sons’ Day
BY D. LYN HUNTER
 |
|
| Caitlin Perry and Nathaniel Bolt take a closer look at the life sciences workshop. |
Is being a scientist like being a big kid who gets to play around with stuff? Do your friends like science too? What do you wear when you’re doing your experiments?
The children packed in the Building 50 auditorium last week peppered Lab researcher Julia Chamberlain with these and many other questions following her brief talk on using bacteria to help clean up radioactive waste.
The occasion was Daughters and Sons to Work Day, an annual event that brings the children of Lab employees to the Hill to learn more about the kind of research that happens here. Last Thursday, the Lab hosted 80 girls and 53 boys, ranging in age from about 9 to 16.
“I am always struck by both the children’s and parents’ anticipation and excitement when they arrive for Daughters and Sons to Work Day,” said Rollie Otto, who heads the Center for Science and Engineering Education which sponsored the event.
After the opening presentation, the children attended several science workshops, learning about such topics as robots, nuclear science, micromachines, liquid nitrogen, and Mars.
Children toured Life Sciences Division labs and then prepared their
own specimens, which they viewed under microscopes. They looked at
their own cheek cells, dissected petals, mounted live maggots, saw
Drosophila embryo genes, and a drop of blood.
David Knowles, who instructed the class with Soile Keranen and Cris
Luengo, said the enthusiasm for learning was extremely high, and even
attracted a visit from Division Director Joe Gray and Deputy Director
Damir Sudar, who helped the kids with their bioimaging.
The day’s activities also included two career fairs —
one for boys and one for girls.
More than 50 Lab employees volunteered for the event — running
workshops, presenting at the career fair, and serving as chaperones.
Among them was Anytra Henderson, an administrative professional who has worked in the Physics Division for the last four years. In addition to serving as a chaperone, Henderson also brought five members of her family, including her niece, nephew, sister, cousin and goddaughter.
“I like bringing them here because they can get hands-on instruction in many areas of science, something they don’t really get in school,” Henderson said. “And they just love it. They look forward to it every year.”
Henderson hopes the visit inspires her relatives to go to college and pursue a career in science. And that, says Otto, is the goal he and other event organizers hope to achieve.